Inter-genomic DNA exchanges and homoeologous gene silencing shaped the nascent allopolyploid coffee genome (Coffea arabica L.)
|
|
|
- Kenneth Miles
- 5 years ago
- Views:
Transcription
1 G3: Genes Genomes Genetics Early Online, published on July 20, 2016 as doi: /g Inter-genomic DNA exchanges and homoeologous gene silencing shaped the nascent allopolyploid coffee genome (Coffea arabica L.) Philippe Lashermes *, Yann Hueber *,1, Marie-Christine Combes *, Dany Severac, Alexis Dereeper * Institut de Recherche pour le Développement, UMR DIADE (IRD, Université de Montpellier), BP 64501, Montpellier Cedex 5, France MGX-Montpellier GenomiX, Institut de Génomique Fonctionnelle, Montpellier Cedex 5, France Institut de Recherche pour le Développement, UMR IPME (IRD, CIRAD, Université de Montpellier), BP 64501, Montpellier Cedex 5, France 1 Present address: BioVersity International, Parc Scientifique Agropolis II, Montpellier Cedex 5, France The entire sequence dataset has been deposited at European Nucleotide Archive under the study accession numbers PRJEB5543 and PRJEB9368 for RNA-seq and DNA-seq, respectively. 1 The Author(s) Published by the Genetics Society of America.
2 Short running title: Genome evolution in Coffea arabica Corresponding author's name: Philippe Lashermes, Institut de Recherche pour le Développement, UMR DIADE, 911 avenue Agropolis, BP 64501, Montpellier Cedex 5, France tel: Key words: polyploidy, evolution, gene conversion, homoeologous recombination, genome dominance. Abstract Allopolyploidization is a biological process that has played a major role in plant speciation and evolution. Genomic changes are common consequences of polyploidization, but their dynamics over time are still poorly understood. Coffea arabica, a recently formed allotetraploid, was chosen to study genetic changes that accompany allopolyploid formation. Both RNA-seq and DNA-seq data were generated from two C. arabica genetically distant accessions. Genomic structural variation was investigated using C. canephora, one of its diploid progenitors, as reference genome. The fate of 9,047 duplicate homoeologous genes was inferred and compared between the accessions. The pattern of SNP density along the reference genome was consistent with the allopolyploid structure. Large genomic duplications or deletions were not detected. Two homoeologous copies were retained and expressed in 96% of the genes analyzed. Nevertheless, duplicated genes were found to be affected by various genomic changes leading to homoeolog loss or silencing. Genetic and epigenetic changes were evidenced that could have played a major role in the stabilization of the unique ancestral allotetraploid and its subsequent diversification. While the early evolution of C. arabica mainly involved homoeologous crossover exchanges, the later stage appears to have relied on more gradual evolution involving gene conversion and homoeolog silencing. 2
3 Introduction Polyploidy (the complete doubling of a genome) has long been recognized as an important mechanism in plant speciation and genome evolution (Soltis et al., 2014). It is now well established that polyploidy occurred frequently during angiosperm evolution and that all flowering plant species have undergone one or more rounds of genome duplication in their history (Jiao et al., 2011). In particular, allopolyploidization (arising from interspecific hybridization accompanied by whole-genome duplication) is considered to have contributed to the adaptation to broader and novel environmental niches and is thought to play a fundamental role in the evolutionary history of speciation (Comai, 2005; Soltis et al., 2014). In addition, many major agricultural crop plants including wheat (Triticum aestivum), cotton (Gossypium hirsutum), rapeseed (Brassica napus), sugarcane (Saccharum officinarum) and coffee (Coffea arabica) are allopolyploids. The establishment of a new allopolyploid species is not a trivial feat. The merging of two or more divergent genomes, and the presence of these parental genomes in duplicate in a single nucleus, can set the stage for dynamic changes to the genome, transcriptome, and phenotype of the new polyploid species (Doyle et al., 2008). However, the newly formed allopolyploid faces several immediate challenges. It must secure exclusive intra-genomic pairing at meiosis that will lead to full fertility and disomic inheritance. Meiosis can therefore have a dual impact on the evolution of many newly formed polyploids by (i) enabling sexual propagation and (ii) generating, through meiotic errors, large-scale chromosomal variation upon which genetic drift and/or selection can act (Leitch & Leitch, 2008). In addition, the newly formed allopolyploid must orchestrate inter-genomic interactions and regulate gene expression to adapt to its environment (Jackson & Chen, 2010). Hence, successful allopolyploidizations are those that trigger an array of genomic changes that confer evolutionary advantages (Arrigo & Barker, 2012). In the last decade, molecular data from resynthesized and natural allopolyploids has shown that genetic and epigenetic changes are common consequences of polyploidization across a wide range of species (Madlung & Wendel, 2013). Nevertheless, little is known about the mechanisms that lead to these changes and even less about their directed or random nature. In addition, genomic responses to whole genome duplication (WGD) appear to be extremely diverse, and both genomic rearrangements and gene expression changes vary to different degrees depending on the polyploid species concerned, thus preventing simple generalizations. For example, rapid genomic changes have been reported in many new 3
4 allopolyploids (Doyle et al., 2008), but not in allopolyploid cotton (Liu et al., 2001). Some polyploid genomes appear to undergo rapid homeolog loss (Gaeta et al., 2007; Soltis et al., 2010; Buggs et al., 2012), whereas in other polyploids, changes in gene expression appear to dominate (Lee & Chen, 2001; Wang et al., 2006; Flagel & Wendel, 2010; Wang et al., 2016). To understand allopolyploid genome evolution in a broad context, genomic data from many more allopolyploids are required. Among allopolyploid plants, Coffea arabica is an interesting case (Lashermes et al., 1999; 2000). C. arabica is a recent (less than years ago) allotetraploid (C a E a genome) formed by hybridization between two diploid species: C. canephora (C genome) and C. eugenioides (E genome). The two parental species are closely related, and the two subgenomes have low sequence divergence (i.e. 1.3% average difference for genes, Cenci et al., 2012). In spite of the close relationship between the two constitutive sub-genomes, C. arabica displays diploid-like meiotic behavior with bivalent formation. In addition, the most recent molecular analyses (Lashermes et al., 2014) showed that genomic rearrangements involving homoeologous exchanges occurred in C. arabica and could be a major source of genetic diversity. Evidence for a large number of homoeologous exchange events (HEEs) shared by all accessions of C. arabica strongly supports the hypothesis of a single allopolyploidization event. The highlands of south west Ethiopia are considered as the primary centre of genetic diversity of C. arabica (Sylvain, 1955). Presence of wild populations has been also reported in forest on the Boma Plateau and Mount Imantong of Sudan (Thomas, 1942) and in Mount Marsabit of Kenya (Anthony et al., 1987). Furthermore, although C. arabica accessions exhibit very low genetic diversity (as estimated through molecular markers), they display marked phenotypic/adaptive variation and there is strong evidence of hybrid vigor when particular accessions are crossed (Van der Vossen et al., 2015). In the present study, C. arabica was used to investigate genomic changes in allopolyploid using high-throughput sequencing approaches. Both RNA-seq and DNA-seq data were generated from two genetically distant accessions, and analyzed using C. canephora as reference genome (Denoeud et al., 2014). On one hand, genomic structural variation was investigated based on mapping patterns of DNA sequence reads and on the detection of copy number alterations (CNA). On the other hand, genes affected by homoeolog loss or silencing were inferred by comparing the number of SNPs detected in C. arabica and between its two diploid progenitor species at both DNA and RNA level. To validate these results, Sanger direct sequencing of cdna and DNA amplicons from various C. arabica 4
5 accessions were performed. In particular, the distribution of two homoeologous crossover exchange events among accessions from the C. arabica primary centre of diversity was investigated. Genomic rearrangements involving homoeologous DNA exchanges as well as gene conversion and homoeolog silencing were evidenced that could have played a major role in the evolution and diversification of C. arabica. Materials and Methods Plant material, library construction and sequencing The plant material came from two C. arabica accessions including the commercial cultivar Caturra (an inbred line) and one wild accession (AR41) from the ORSTOM (1966) collection mission in Ethiopia, and one accession from each of the two modern-day diploid progenitors, C. canephora (acc. DH200-94) and C. eugenioides (acc. DA58). Young leaf tissues were collected at the same time from individuals plants grown in a greenhouse in Montpellier (France) and were immediately flash frozen in liquid nitrogen and stored at -80 C until DNA and RNA extractions as previously reported (Combes et al., 2013; Lashermes et al., 2014). For genomic DNA sequencing of the two C. arabica accessions, non-indexed paired-ends (PE) libraries were constructed using the TruSeq DNA sample preparation kit (Illumina), which includes a fragmentation step by sonication and after end-repair and adapter ligation, selection of 470 ± 60 bp DNA fragments by band excision after gel electrophoresis. Pairedend sequencing was carried out at 2 x 100 bp using the Illumina HiSeq 2,000 according to the manufacturer s instructions at the MGX platform (Montpellier Genomix, France). DNA data were collected from two lanes (one lane per library and per accession) in the same sequencing run. For RNA sequencing of C. arabica and C. eugenioides accessions, the method and data were as previously reported in (Lashermes et al., 2014). Briefly, total RNA was isolated from 1 g of material from each plant and mrna libraries were constructed using the Illumina RNA-seq sample prep kit (Illumina, San Diego, CA, USA). Single-reads (~72 nt) were generated with the Illumina HiSeq 2,000. For the accession Caturra (C. arabica), 4 independent libraries were constructed from 4 different plants. The entire sequence dataset has been deposited at European Nucleotide Archive under the study accession numbers PRJEB5543 and PRJEB9368 for RNA-seq and DNA-seq, respectively. Analysis of DNA-seq data 5
6 Sequences were first cleaned to remove adapter sequences and quality filtered (phred score higher than 28). After trimming, reads of less than 50 bp were discarded. Paired-end sequences were then mapped onto the total C. canephora (acc. DH200-94) reference genome ( Denoeud et al., 2014) using the BWA-MEM algorithm (Li & Durbin, 2010; with the paired-end mode and the default parameters. The resulting BAM files of unambiguously aligned sequences were then analyzed for SNP discovery with the GATK toolkit ( using the Unified-Genotyper module with default parameters. For the detection of copy number changes, the BAM files were analyzed with FREEC (control-free Copy number caller, Boeva et al., 2011) using a non-overlapping 50 kb sliding window. The main steps are (i) normalization of the copy number profiles using GC content, (ii) segmentation of normalized profiles and (iii) assignment of copy number changes to losses and gains. SNP density was estimated along the 11 homoeologous chromosome groups and regions exhibiting homoeologous SNP deficit (HSD) were identified using in-house Perlscripts (available upon request). Non-overlapping 10 kb sliding windows were used to estimate the SNP density along the C. canephora reference genome. To minimize the rate of false-positive SNPs, a minimum depth coverage of 10 was required for a position to be considered while positions exhibiting depth coverage greater than twice the overall sample mean were discarded. Since the genome sequence of C. eugenioides is not yet available, expected local SNP density (in the absence of genetic changes in C. arabica) along the homoeologous chromosome groups could not be determined. So, the overall mean SNP density was used as reference value, and the hypothesis of HSD was retained when at least five consecutive 10 kb windows (presenting a minimum of 8,000 positions that satisfied the depth criteria) along the C. canephora reference genome were identified as exhibiting a highly significant SNP deficit using a test of proportions with R software (prop.test; Newcombe, 1998). To control for multiple testing, the resulting P value were corrected using the Benjamini and Hochberg procedure (Benjamini & Hochberg, 1995). A cumulative window size of 50 kb was found to produce a good compromise between informativeness and accuracy. RNA-seq data processing The 72-nt reads of each library were mapped to a C. canephora coding DNA sequence reference (25574 CDS) as transcriptome reference using BWA MEM with the default parameters ( Denoeud et al., 2014). The aligned sequences of each 6
7 library were then analyzed to find SNPs (single nucleotide polymorphisms) with the GATK toolkit using the Unified-Genotyper module with default parameters to obtain a list of SNPs and allelic data, and the Depth-Of-Coverage module to obtain information on depth coverage. Regarding the accession Caturra, analyses were performed using either a pool of reads from the four libraries or each individual library. To avoid artefacts due to reads from pseudogenes or repeat sequences, only CDS identified as single copy were used for subsequent analyses. Comparison of SNPs determined by RNA-seq and DNA-seq SNPs were quantified and compared using SNiPlay ( a dedicated web-based tool (Dereeper et al., 2011). In particular, SNiPlay authorizes the user to set minimum depth coverage for a sequence position to be taken into consideration. To allow combined analysis of SNP-gene data generated by both RNA-seq and DNA-seq, the list of SNPs, allelic data, and depth coverage corresponding to the CDS were extracted from the whole genome DNA-seq data using the in-house Perl-script (available upon request) and the GFF file of the C. canephora reference genome ( For each gene, the SNPs detected in each of accessions analyzed by either DNA-seq or RNA-seq were determined and common sequence positions were compared. A minimum depth coverage of 12 was required for the DNA-seq in C. arabica to ensure a good coverage of the two subgenomes while the RNA-seq read cutoff was set at 4 and 16 for C. eugenioides and C. arabica, respectively, to account for the possibility of SNP allele low frequency due to homoeolog expression bias in the allopolyploid (Yoo et al., 2014; Combes et al., 2013). In addition, SNP cut-off parameters were applied throughout the analyses in order to take into account the occurrence of sequencing (or SNP detection) errors and the fact that the two diploid parental accessions used in this experiment are not the true diploid progenitors of the allotetraploid C. arabica. The fate of homoeologous genes in C. arabica was investigated as shown in Figure 1. For each gene, SNP number and DNA read depth coverage comparisons were carried out. A minimum of three SNPs between the two diploid species as well as in the allopolyploid as determined by DNA-seq were selected as the cutoff to set a lower bound on the resolution of SNP number change for a given gene. First, the number of SNPs determined by DNA-seq in the allopolyploid (SNP4X-D) was compared to the number of SNPs between the two diploid species (SNP2X). For a given gene, when SNP2X and SNP4X-D were comparable, two homoeologous copies were considered to have been retained. In contrast, genes displaying at least a four-fold decrease between SNP2X and SNP4X-D, and no more than one SNP 7
8 classified as homoeologous (SNP4X-D 1) were considered as exhibiting homoeolog loss in the C. arabica accession concerned (when SNP4X-D was null, a minimum value of 3 was required for SNP2X). Second, for each gene presenting two homoeologous copies, the number of SNPs determined in the allopolyploid by RNA-seq (SNP4X-R) and DNA-seq (SNP4X-D) were compared. Genes in the allopolyploid displaying at least three SNPs at the DNA level (SNP4X-D 3) and no SNPs at the RNA level (SNP4X-R = 0; presence of one SNP was tolerated if different than those detected by DNA-seq) were considered as exhibiting homoeolog silencing in the C. arabica accession concerned. Others genes displaying similar SNPs at the DNA and RNA levels were considered expressing both homoeologs. Finally, for each gene presenting one homoeolog loss, the average read coverage was compared to the expected values (99% confidence interval) for a gene with two copies and four copies, respectively, as reported previously (Lashermes et al., 2014). According to the gene read depth coverage categorization, the homoeolog loss was inferred to result from either sequence deletion (i.e. two copies) or sequence homogenization (i.e. four copies). Analysis of genes exhibiting homoeolog loss and homoeolog silencing In each gene involved in homoeolog loss and homoeolog silencing, the numbers of SNPs shared by C. arabica and either C. canephora or C. eugenioides were compared to identify the subgenome in which the event occurred. Homoeolog loss/silencing was attributed to the subgenome deriving from the diploid progenitor species with the smallest number of SNPs shared with C. arabica. A difference between the two SNP categories of at least three SNPs was required to determine the subgenome in which the homoeolog loss or silencing occurred. Computational mapping and Plant GO-slim annotation were performed using Blast2GO software v2.6.4 ( Conesa & Gotz, 2008) as described in Combes et al. (2013). Functional enrichments in groups of interesting genes were investigated using Fisher s exact test, applying a false discovery rate (FDR), and a correction for multiple testing. Validation of genes exhibiting homoeolog lost and silencing Experimental validation of bioinformatically inferred genes exhibiting either homoeolog lost or silencing was performed using traditional Sanger sequencing as previously reported in Combes et al. (2012). For homoeolog lost analysis, three genes from the two main regions (i.e. B and C, see Table 2) showing contiguous genes exhibiting homoeolog losses in both accessions analyzed were selected (Supporting information Table S4). Primer pairs were 8
9 designed to amplify DNA fragments containing species-specific SNPs that differentiate the two diploid progenitor species, C. canephora and C. eugenioides (Combes et al., 2015). SNP detection assays were performed based on Sanger direct sequencing of DNA amplicons from 96 accessions originating from the main regions of C. arabica primary centre of diversity (Supporting information Table S5). For silencing validation, a subset of 5 genes was selected and primer pairs were designed to amplify single-exon fragments containing several homoeologous SNP (Supporting information Table S4). The expression of homeologs was analysed using a SNP ratio quantification method based on dideoxy-terminated sequences of cdna and DNA amplicons from C. arabica (acc. Caturra, AR41 and AR59) as described in Combes et al. (2012). PCR reactions were performed twice in a volume of 25 µl with either 1 µl of the diluted (one-tenth) cdna generated by the first-strand synthesis or 25 ng of genomic DNA, 0.4 µm of each primer, 2.5 mm MgCl2, and 1 U of Taq DNA Polymerase. Cycling was done in a GeneAmp PCR 9700 thermocycler for 2 min at 94 C followed by 5 cycles of 10 sec at 94 C, 30 sec at 60 C to 55 C (-1 C/cycle) and by 30 cycles of 10 sec at 94 C, 30 sec at 55 C, 30 sec at 72 C, and a final 8-min extension at 72 C. Sequencing was performed by Beckman Coulter Genomics (Takeley UK). Sequence chromatograms of exon portions amplified on cdna and DNA were analysed and compared using BioEdit (Hall, 1999). Results Genomic structural variation Genome sequencing of the two genetically distant accessions of C. arabica yielded 286 million 100-bp paired-end reads comprising 57.3 Gb of raw data (Supporting information Table S1). After cleaning, 267 million (i.e. 93%) of the reads remained and were mapped to the high-quality draft C. canephora genome sequence. The number of reads mapped onto the genome ranged from 106 million in accession AR41 to 159 million in accession Caturra. The average sequence depth of coverage of the reference canephora genome varied from 36x to 51x depending on the C. arabica accession. Copy number alteration (CNA) based on the depth of coverage was assessed using FREEC (Boeva et al., 2011). Copy number profiles were established for the two C. arabica accessions using a non-overlapping 50 kb sliding window and normalization of GC content. The outcome for the accession Caturra is shown in Figure 2 as an example. Although few CNAs were predicted for each accession, the overall profiles appeared to be consistent with a 4X copy number along the 11 C. canephora chromosomes used as reference. Gross copy 9
10 number changes due to large genomic duplications or deletions were not detected. These results are coherent with the expected allotetraploid structure of C. arabica. SNPs were detected in each of the two accessions of C. arabica. In allopolyploids such as C. arabica, sequence variations between subgenomes (homoeologous SNPs) co-exist with allelic variations within subgenomes (homologous SNPs). However, given the high level of homozygosity in C. arabica (Lashermes et al., 2014), it was assumed that most of the identified SNPs are homoeologous SNPs. Non-overlapping 10 kb sliding windows and coverage criteria required to consider a position to minimize the rate of false SNPs were applied to estimate the density of SNPs along the 11 homoeologous chromosome groups. Whatever the accession considered, the overall SNP density appeared to be relatively constant along the different reference chromosomes, with a slight increase in the genomic regions enriched in transposable elements (Figure 3). In the two accessions and the regions analyzed, the structural heterozygosity observed was in line with what is expected in an allotetraploid. Nevertheless, regions exhibiting homoeologous SNP deficit (HSD) were identified (Figure 4), for example, 39 regions exhibiting HSD were revealed in C. arabica acc. Caturra (Supporting information Table S2). Of these regions, 37 (i.e. 95%) were shared by the two accessions analyzed. In addition, except the region (1,170 kb in size) identified on chromosome 7, the regions were rather small, ranging from 50 to 160 kb with an average of 81 kb. The overall size of these regions was 4,230 kb, representing 1.5% of the genome windows analyzed that satisfied the depth criteria. Evidence of homoeolog loss The fate of homoeologous genes in C. arabica was inferred as described in Figure 1. The number of SNPs per gene detected by DNA-seq in individual accessions of C. arabica were compared to those detected between accessions of its two diploid progenitor species in common sequence positions. A set of 9,047 genes that satisfied depth and quality requirements was analyzed in the two arabica accessions. While two homoeologous copies appeared to be retained in 98% of the genes, 179 genes (2.0%) exhibiting homoeolog loss were identified (Table 1). Although the number of homoeolog losses varied from 148 (Caturra) to 174 (AR41) among the two C. arabica accessions, a large proportion (80%) of genes exhibiting homoeolog loss was shared by the two accessions. The distribution across the reference genome of loci exhibiting homoeolog loss was next investigated (Figure 5). While homoeolog loss events were observed across the 11 reference chromosomes, four regions carrying contiguous genes exhibiting homoeolog losses 10
11 were detected (Table 2). The number of genes per region in the accession Caturra ranged from three to 142, and the corresponding genome fragment size ranged from 15 to 1,198 kb. The four regions were shared by the two accessions and corresponded to genomic regions previously identified as exhibiting HSD. In contrast, all the single genes exhibiting homoeolog losses that did not belong to the four identified clusters corresponded to genome regions (10 kb windows) exhibiting standard SNP density. The subgenome origin of the detected homoeolog loss events was further investigated (Table 1). Homoeolog loss was attributed to the subgenome deriving from the diploid progenitor species with the least SNPs shared with C. arabica. Whatever the accession, homoeolog losses were attributed to the two subgenomes, but with a marked preference (at least 75%) for subgenome C a. Contrasted behavior was observed between the groups of genes exhibiting homoeolog loss shared by the two accessions or restricted to one accession. While the homoeolog losses shared by accessions were mainly attributed to subgenome C a (i.e. 81 %), the subgenome origin of homoeolog loss events specific to one accession appeared to be balanced between C a and E a. The mechanisms behind the homoeolog loss events were further investigated. To distinguish between sequence deletion and sequence homogenization, the gene copy numbers in C. arabica were estimated using the DNA-seq read depth of coverage (Figure S1). Indeed, homoeolog sequence deletion in an allotetraploid is expected to be associated with a decrease in gene copy number from 4X to 2X. The distribution of genes exhibiting homoeolog loss according to their average read depth was very similar to the distribution of the overall genes supporting the hypothesis that most of the observed homoeolog losses are not associated with mere gene loss. Nevertheless, genes that exhibited an average read depth corresponding to the expected coverage for gene in two copies were overrepresented in the group of genes exhibiting homoeolog loss (Chi-squared test, p-value < ) suggesting that a few homoeolog loss events could result from sequence deletion. For instance in C. arabica acc. Caturra, of the 148 genes identified as exhibiting homoeolog loss, 12 (8.1%) displayed an average read depth corresponding to the expected coverage for gene in two copies. Homoeolog loss was validated using direct Sanger sequencing of amplicons for three genes (Supporting information Table S4) from the two main regions showing contiguous genes exhibiting homoeolog losses in the two examined arabica accessions (i.e. regions B and C, see Table 2). The frequency of these homoeolog loss events among the C. arabica germplasm was further investigated. These events appeared shared by all of the 96 analyzed 11
12 accessions that represent the main coffee growing regions in the C. arabica primary centre of diversity. Evidence of homoeolog silencing The occurrence of homoeolog silencing in C. arabica accessions was inferred by comparing the number of SNPs determined by DNA-seq (SNP4X-D) and RNA-seq (SNP4X-R) in common sequence positions in each individual gene (Figure 1). It was assumed that in the absence of homoeolog silencing in C. arabica, the number of SNPs determined by DNA-seq and RNA-seq would be equivalent, whereas the failure to detect homoeologous SNPs by RNA-seq would reveal homoeolog silencing. For most of the 9,047 genes analyzed in the two accessions, the number of SNPs determined by DNA-seq and RNA-seq was comparable and overall, 95% of the SNPs identified with the two methods were similar. Taking into account the two C. arabica accessions, 156 genes (1.7%) exhibiting homoeolog silencing were identified (Table 1). The number of genes exhibiting homoeolog silencing varied from 116 (Caturra) to 120 (AR41) between the two accessions of C. arabica, with 80 genes (51.3%) shared by the two accessions and 49% of the silenced genes specific to one of the accessions. Homoeolog silencing were examined among the four individual plants of the accession Caturra. After filtering for the depth coverage, 66 of 120 genes exhibiting homoeolog silencing in Caturra were investigated. For all of them, homoeolog silencing was observed in all individual plants. The distribution across the reference genome of loci exhibiting homoeolog silencing was next investigated. The distribution observed for accession Caturra is shown in Figure 5 as an example. Homoeolog silencing events were observed across the 11 reference chromosomes, and two small regions (92 and 59 kb respectively on chromosome 2 and unanchored scaffold) carrying a cluster of genes exhibiting homoeolog silencing were detected in two accessions. The subgenome origin of the homoeolog silencing events was further investigated (Table 1). Homoeolog silencing was attributed to the subgenome deriving from the diploid progenitor species displaying the smallest number of SNPs shared with C. arabica. Whatever the accession, homoeolog silencing was attributed to the two subgenomes with a marked preference for subgenome C a (from 60.6% to 62.2% depending on the accession). This preference (i.e. 66.2% of C a silenced homoeologs) was particularly observed among the group of genes exhibiting homoeolog silencing shared by the two accessions. 12
13 Homoeolog silencing was validated using direct Sanger sequencing of cdna and DNA amplicons from C. arabica (acc. Caturra, AR41 and AR59). Primer pairs were designed to amplify single-exon fragments of 5 genes for which homoeolog silencing was bioinformatically inferred (Supporting information Table S2). For all the genes assayed, expressions of only one homoeolog from either the subgenomes C a or E a were detected as expected from the previous analyses. Gene ontology enrichment analysis Gene ontology enrichment was analyzed in groups of genes showing either homoeolog loss or homoeolog silencing using the full set of analyzed genes as reference group. None significant variation was observed in the group of genes exhibiting homoeolog loss, whereas statistically highly significant impoverishments in very general GO terms were observed in the group of genes exhibiting homoeolog silencing (Supporting information Table S3). In particular, specific functions related to macromolecular biosynthesis and organic cyclic compound metabolism were under-represented in the group of genes exhibiting homoeolog silencing shared by the two accessions (Table 3). Discussion Plant allopolyploidy appeared to be associated with an array of rapid genomic changes in genetic/epigenetic, transcriptomic and proteomic layers that may affect the fitness of the newly formed allopolyploid and increase its competitiveness, leading to its successful establishment in nature (Arrigo & Barker, 2012). In the present study, several approaches based on high-throughput sequencing technologies were used to investigate these genetic changes in C. arabica, a model allopolyploid perennial plant. The use of a high-quality draft genome sequence of C. canephora (Denoeud et al., 2014) as genome and transcriptome references offered new opportunities for analyses compared with previous works (Lashermes et al., 2014). However, inherent limitations remained such as the impossibility to study putative genomic regions of C. arabica that do not have a counterpart in the C. canephora reference genome, and the fact that interpretation of the subgenome (i.e. parental) of the identified homoeologous SNPs was limited to the part of the genome for which reference sequences of both diploid parents were available. Nevertheless, whole genome analyses and comparison of two geographically and genetically distant accessions of C. arabica provided 13
14 compelling evidence for the efficiency of sequencing based methods for the investigation of genetic changes in allopolyploid and of the mechanisms that lead to these changes, their timing, and their directed or random nature. Fixed heterozygosity and allopolyploid structure Although challenging because of the low divergence between the two diploid constitutive sub-genomes, the allotetraploid genome organization of C. arabica was first revealed by genomic in situ hybridization (Lashermes et al., 1999). The present data confirm a state of fixed heterozygosity related to the presence of two complete sets of homoeologous chromosomes in C. arabica. Whatever the accession considered, the observed pattern of SNP density along the 11 homoeologous chromosome groups in C. arabica is consistent with its assumed allopolyploid structure. Furthermore, large genomic duplications or deletions were not detected, confirming low chromosomal divergence between genomes E and C of the two diploid progenitor species (Pinto-Maglio, 2006) and suggesting that, following the original hybridization event, overall genome organization is stable in C. arabica. Evidence for homoeologous exchanges Homoeologous exchanges appear to have played an important role in the evolution of the C. arabica genome. Indeed, regions exhibiting homoeologous SNP deficit (HSD) and 4X copy number were identified across the genome. These regions, which ranged from 50 kb to 1,170 kb in size, suggest major genetic changes. Although certainly underestimated, since the method used here cannot resolve small regions (a minimum of 50 kb was required), these regions represented nearly 1.5% of the portion of C. arabica genome analyzed. Furthermore, among the 9,047 duplicate homoeologous genes whose fate was successfully inferred in the two accessions of C. arabica analyzed, 2.0% were found to be affected by genomic changes leading to homoeolog loss without sequence deletion in most cases sequence. Two mechanisms have been suggested to lead to such exchanges of homoeologous chromatids: via crossovers and the subsequent segregation of one parental and one recombinant chromatid, or via non-crossover exchange, also called gene conversion (Gaeta & Pires, 2010). Due to their large size, homoeologous crossover exchanges are the most likely mechanism at the origin of the C. arabica regions exhibiting HSD. In contrast, both mechanisms are hypothesized to be involved in the occurrence of genes exhibiting homoeolog losses. While the regions carrying contiguous genes exhibiting homoeolog losses correspond to regions exhibiting HSD, most single homoeolog loss events likely result from gene conversion. The high frequency of 14
15 homoeologous contact across the 11 homoeologous chromosome groups is a surprising result given the apparently complete bivalent formation at meiosis. However, meiotic abnormalities have been repeatedly observed in C. arabica (Grassias & Kammacher, 1975; Owuor, 1985). Although exceptional, weakness in the control of the chromosome pairing in C. arabica could therefore enable homoeologous exchanges. A possibility is that most of these homoeologous exchanges, having occurred early, could be the result of possible multivalent (or less strict bivalent) pairing at meiosis at inception. Similar high frequencies of non-crossover homoeologous exchanges have been reported in other genome-wide analyzed allotetraploids such as Gossypium hirsutum and Brassica napus (Salmon et al., 2010; Guo et al., 2014; Chalhoub et al., 2014) suggesting an important mechanism by which allopolyploid genomes adapt to the duplicate state. Furthermore, the two main homoeologous crossover exchanges detected in the two sequenced accessions of C. arabica were also evidenced in all analysed accessions (i.e. 96 accessions) originating from the main regions of C. arabica primary centre of diversity. This observation strongly support the hypothesis of a single origin for C. arabica (Lashermes et al. 2014). All C. arabica populations would derive from a unique allopolyploidization event associated with large and specific homoeologous crossover exchanges. Homoeolog silencing suggests epigenetic changes Gene silencing is a common response to polyploidy and has been described in many allopolyploids, including Arabidopsis (Wang et al., 2006), cotton (Chaudhary et al., 2009), Tragopogon (Buggs et al., 2011) and wheat (Bottley et al., 2006). Silencing can occur as early as the first generation following polyploidy and some duplicated genes may be silenced in some organs of the plant but continue to be expressed in other organs (Adams & Wendel, 2005). By combining RNA-seq and DNA-seq data, homoeolog silencing was inferred for 1.7% of genes in the two accessions of C. arabica analyzed in the present study. Since gene expression was estimated only in leaves, partition of the expression of duplicate genes to specialized tissue-specific expression activity was not investigated. Silencing mechanisms almost certainly vary with the gene. In the absence of sufficient time for point mutations to accumulate in promoter regions or other cis-regulatory elements, most gene silencing is believed to be epigenetically induced (Adams & Wendel, 2005). Intergenomic interactions between progenitor genomes in allopolyploids are predicted to induce epigenetic changes including histone modifications and DNA methylation (Song & Chen, 2015). In the last few years, gene silencing via epigenetic changes has been documented in several plants including 15
16 Tragopogon (Sehrish et al., 2014), Arabidopsis (Chen et al., 2008) and Brassica rapa (Cheng et al., 2016). Two temporally distinct phases of evolution The implication of homoeolog loss and silencing events in the establishment and diversification of C. arabica is questionable (Madlung & Wendel, 2013). Comparison of the two analyzed accessions revealed the occurrence of shared genomic changes, whereas other events appeared specific to one accession. While 80% of genes exhibiting homoeolog loss were shared by the two accessions, only half of genes exhibiting homoeolog silencing seemed shared by the two accessions. In particular, the four detected genomic regions exhibiting HSD and contiguous homoeolog losses were shared by the two accessions. Two temporally distinct phases of evolution can therefore be hypothesized. A first phase accompanying the allopolyploidization process and mainly involving homoeologous crossover exchanges could have been followed by a more gradual phase of duplicate gene evolution involving gene conversion and homoeolog silencing. Furthermore, patterns of gene loss and retention could be explained by the genebalance hypothesis (Freeling, 2009; Feldman et al., 2012). Under this hypothesis, the evolution of genes is linked to their function within networks (Edger & Pires, 2009), so genes coding for products that are closely connected would be dosage sensitive genes and are thought to be retained or eliminated together to preserve stoichiometry. This evolution mechanism was recently proposed for homoeologs in T. miscellus, a young natural allopolyploid species (Buggs et al., 2012). This process is probably marginal in the pattern of gene loss observed in C. arabica given the lack of significant GO term enrichment in the subset of genes showing homoeolog loss. However, an equivalent trend might play a role in the gene silencing pattern of C. arabica. Indeed, genes related to macromolecular biosynthesis and organic cyclic compound metabolism tended to be preserved from gene silencing, suggesting gene dosage requirements (Veitia et al., 2013). Asymmetric evolution Studies on duplicated genomes or genomic regions in ancient polyploids showed that they have often experienced unequal gene losses (or genome fractionation), with one genome or genomic region retaining more genes (dominant) than the other (more fractionated). More interestingly, genes located on the dominant genome or genomic region tend to have higher expression levels (Schnable et al., 2011; Cheng et al., 2012; Garsmeur et al., 2014; Li et al. 16
17 2014). Such genome dominance phenomena have been reported in a few plants including Arabidopsis thaliana (Wang et al., 2006), Brassica (Cheng et al., 2012), maize (Schnable et al., 2011) and wheat (Li et al. 2014; Wang et al., 2016). In C. arabica, homoeolog loss and silencing were attributed to both subgenomes with a marked preference for subgenome C a, suggesting dominance of the E a genome. However, neither of the two subgenomes appeared to be preferentially expressed in C. arabica (Combes et al., 2013) or in interspecific hybrids between C. canephora and C. eugenioides (Combes et al., 2015). The absence of relationships between the observed preferential changes in the C a genome and overall homoeologous expression could be linked to the recent origin and the intertwined homoeolog regulations occurring in C. arabica (Combes et al., 2013). Conclusions The present results point to overall structural genome stability in C. arabica following the original hybridization event. However, homoeologous DNA exchanges and homoeolog silencing were evidenced that could have played a major role in the stabilization and survival of the ancestral allotetraploid and in its subsequent diversification. While the early phase of evolution mainly involved homoeologous crossover exchanges, the later phase appears to have relied on a more gradual phase of duplicate gene evolution involving gene conversion and homoeolog silencing. As suggested in C. arabica, the evolutionary success of a newlyformed polyploidy may require a delicate balance between genetic and epigenetic changes. Acknowledgements This work was partially supported by a grant from the Direction de la Valorisation au Sud de l Institut de Recherche pour le Développement (IRD, France). We thank Dr. V. Boeva for advice on the use of the FreeC tool to identify copy number alteration. Also we acknowledge M. S. Seyoum (Rothamsted International African Fellow-2006) from the Awada Coffee Research Centre for providing leaf samples. Supplemental material Table S1. Total number of reads sequenced and mapped onto the C. canephora (acc. DH200-94) reference genome after sequencing of two accessions of C. arabica. 17
18 Table S2. Identification of regions exhibiting putative homoeologous SNP deficit (HSD) in C. arabica (acc. Caturra). Table S3. Gene ontology enrichment analysis of genes exhibiting homoeolog silencing in C. arabica. Table S4. Genes investigated, pairs of oligonucleotides and SNP used to validate genes exhibiting homoeolog lost or silencing in C. arabica. Table S5. List of C. arabica accessions used to validate the presence of genomic regions exhibiting homoeologous SNP deficit and to analyze their distribution among the C. arabica germplasm. Except four varieties, those accessions are local landraces from garden coffee (AWADA collection) or plants from forest (IRD & CATIE collections) collected in the major coffee growing regions of Ethiopia (Labouisse et al. 2008). Figure S1. Read depth measurements of genes of C. arabica (acc. Caturra). Distribution is shown according to the average read depth of coverage of all genes, genes exhibiting homoeolog loss or silencing. Only the set of 9,047 genes analyzed in the two accessions was taken into consideration. References Adams KL, Wendel JF Polyploidy and genome evolution in plants. Current Opinion in Plant Biology 8: Anthony F, Berthaud J, Guillaumet JL, Lourd M Collecting wild Coffea species in Kenya and Tanzania. Plant Genetic Resources Newsletter 69: Arrigo N, Barker MS Rarely successful polyploids and their legacy in plant genomes. Current Opinion in Plant Biology 15: Benjamini Y, Hochberg Y Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society, Series B 57: Boeva V, Zinovyev A, Bleakley K, Vert JP, Janoueix-Lerosey I, Delattre O, Barillot E Control-free calling of copy number alterations in deep-sequencing data using GCcontent normalization. Bioinformatics 27: Bottley A, Xia GM, Koebner RMD Homoeologous gene silencing in hexaploid wheat. Plant Journal 47: Buggs RJA, Chamala S, Wu W, Tate JA, Schnable PS, Soltis DE, Soltis PS, Barbazuk WB Rapid, repeated, and clustered loss of duplicate genes in allopolyploid plant populations of independent origin. Current biology 22: Buggs RJA, Zhang L, Miles N, Tate JA, Gao L, Wu W, Schnable PS, Barbazuk WB, Soltis PS, Soltis DE Transcriptomic shock generates evolutionary novelty in a newly formed, natural allopolyploid plant. Current biology 21:
19 Cenci A, Combes MC, Lashermes P Genome evolution in diploid and tetraploid Coffea species as revealed by comparative analysis of orthologous genome segments. Plant molecular biology 78: Chalhoub B, Denoeud F, Liu S, Parkin IA, Tang H, Wang X, Chiquet J, Belcram H, Tong C, Samans B, Corréa M et al Early allopolyploid evolution in the post- Neolithic Brassica napus oilseed genome. Science 345: Chaudhary B, Flagel L, Stupar RM, Udall J, Verma N, Springer NM, Wendel JF Reciprocal silencing, transcriptional bias and functional divergence of homeologs in polyploid cotton (Gossypium). Genetics 182: Chen M, Ha M, Lackey E, Wang J, Chen ZJ RNAi of met1 reduces DNA methylation and induces genome-specific changes in gene expression and centromeric small RNA accumulation in Arabidopsis allopolyploids. Genetics 178: Cheng F, Wu J, Fang L, Sun S, Liu B, et al Biased gene fractionation and dominant gene expression among the subgenomes of Brassica rapa. Plos One 7:e36442 Cheng F, Sun C, Wu J, Schnable J, Woodhouse MR, Liang J, Cai C, Freeling M, Wang X Epigenetic regulation of subgenome dominance following whole genome triplication in Brassica rapa. New Phytologist doi: /nph Comai L The advantages and disadvantages of being polyploid. Nature Reviews Genetics 6: Combes MC, Cenci A, Baraille H, Bertrand B, Lashermes P Homeologous gene expression in response to growing temperature in a recent allopolyploid (Coffea arabica L.). Journal of Heredity 103: Combes MC, Dereeper A, Severac D, Bertrand B, Lashermes P Contribution of subgenomes to the transcriptome and their intertwined regulation in the allopolyploid Coffea arabica L. grown at contrasted temperatures. New Phytologist 200: Combes MC, Hueber Y, Dereeper A, Rialle S, Herrera JC, Lashermes P Regulatory divergence between parental alleles determines gene expression patterns in hybrids. Genome Biology and Evolution 7: Conesa A, Gotz S Blast2GO: A comprehensive suite for functional analysis in plant genomics. International Journal of Plant Genomics, ID , Denoeud F, Carretero-Paulet L, Dereeper A, Droc G, Guyot R, Pietrella M, Zheng C, Alberti A, Anthony F, Aprea G et al The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science 345: Dereeper A, Nicolas S, Lecunff L, Bacilieri R, Doligez A, Peros JP, Ruiz M, This P SNiPlay: a web-based tool for detection, management and analysis of SNPs. Application to grapevine diversity projects. BMC Bioinformatics 12(1):134. Doyle JJ, Flagel LE, Paterson AH, Rapp RA, Soltis DE, Soltis PS, Wendel JF Evolutionary genetics of genome merger and doubling in plants. Annual Review of Genetics 42:
20 Edger PP, Pires JC Gene and genome duplications: the impact of dosage-sensitivity on the fate of nuclear genes. Chromosome Research 17: Feldman M, Levy A, Fahima T, Korol A Genomic asymmetry in allopolyploid plants: wheat as a model. Journal of Experimental Botany 63: Flagel LE, Wendel JF Evolutionary rate variation, genomic dominance and duplicate gene expression evolution during allotetraploid cotton speciation. New Phytologist 186: Freeling M Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annual Review of Plant Biology 60: Gaeta RT, Pires JC Homoeologous recombination in allopolyploids: the polyploid ratchet. New Phytologist 186: Garsmeur O, Schnable JC, Almeida A, Jourda C, D Hont A, Freeling M Two evolutionarily distinct classes of paleopolyploidy. Molecular Biology and Evolution 31: Grassias M, Kammacher P Observations sur la conjugaison chromosomique de Coffea arabica L.. Café Cacao Thé 19: Guo H, Wang X, Gundlach H, Mayer KFX, Peterson DG, Scheffler BE, Chee PW, Paterson AH Extensive and biased intergenomic nonreciprocal DNA exchanges shaped a nascent polyploid genome, Gossypium (Cotton). Genetics 197: Hall TA BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium 41: Jackson S, Chen ZJ Genomic and expression plasticity of polyploidy. Current Opinions in Plant Biology 13: Jiao YN, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang HY, Soltis PS et al Ancestral polyploidy in seed plants and angiosperms. Nature 473: Labouisse JP, Bellachew B, Kotecha S, Bertrand B Current status of coffee (Coffea arabica L.) genetic resources in Ethiopia: implications for conservation. Genetic Resources & Crop Evolution 55: Lashermes P, Combes MC, Hueber Y, Severac D, Dereeper A Genome rearrangements derived from homoeologous recombination following allopolyploidy speciation in coffee. Plant Journal 78: Lashermes P, Combes MC, Robert J, Trouslot P, D Hont A, Anthony F, Charrier A Molecular characterisation and origin of the Coffea arabica L. genome. Molecular Genetics and Genomics 261: Lashermes P, Paczek V, Trouslot P, Combes MC, Couturon E, Charrier A Singlelocus inheritance in the allotetraploid Coffea arabica L. and interspecific hybrid C. arabica x C. canephora. Journal of Heredity 91:
21 Lee HS, Chen ZJ Protein-coding genes are epigenetically regulated in Arabidopsis polyploids. Proceedings of the National Academy of Sciences of the United States of America 98: Leitch AR, Leitch IJ Perspective - Genomic plasticity and the diversity of polyploid plants. Science 320: Li H, Durbin R Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinforma 25: Li A, Liu D, Wu J, Zhao X, Hao M, Geng S, Yan J, Jiang X, Zhang L, Wu J, et al mrna and small RNA transcriptomes reveal insights into dynamic homoeolog regulation of allopolyploid heterosis in nascent hexaploid wheat. Plant Cell 26: Liu B, Brubaker CL, Mergeai G, Cronn RC, Wendel JF Polyploid formation in cotton is not accompanied by rapid genomic changes. Genome 44: Madlung A, Wendel JF Genetic and epigenetic aspects of polyploid evolution in plants. Cytogenetic and Genome Research 140: Newcombe RG Two-Sided Confidence Intervals for the Single Proportion: Comparison of Seven Methods. Statistics in Medicine 17: Owuor JBO Interspecific hybridization between Coffea arabica L. and tetraploid C. canephora P. Ex Fr. II. Meiosis in F 1 hybrids and back crosses to C. arabica. Euphytica 34: Pinto-Maglio CAF Cytogenetics of coffee. Brazilian Journal of Plant Physiology 18: Salmon A, Flagel L, Ying B, Udall JA, Wendel JF Homoeologous nonreciprocal recombination in polyploid cotton. New Phytologist 186: Schnable JC, Springer NM, Freeling M Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proceedings of the National Academy of Sciences of the United States of America 108: Sehrish T, Symonds VV, Soltis DE, Soltis PS, Tate JA Gene silencing via DNA methylation in naturally occurring Tragopogon miscellus (Asteraceae) allopolyploids. BMC Genomics 15: 701 Soltis DE, Buggs RJA, Doyle JJ, Soltis PS What we still don t know about polyploidy. Taxon 59: Soltis DE, Visger CJ, Soltis PS The polyploidy revolution then...and now: Stebbins revisited. American Journal of Botany 101: Song Q, Chen ZJ Epigenetic and developmental regulation in plant polyploids. Current Opinion in Plant Biology 24: Sylvain PG Some observations on Coffea arabica L. in Ethiopia. Turrialba 5: Thomas AS The wild arabica coffee on the Boma Plateau, Anglo-Egyptian Sudan. Empire Journal Experimental Agriculture 10:
22 Van der Vossen HAM, Bertrand B, Charrier A Next generation variety development for sustainable production of arabica coffee (Coffea arabica L.): a review. Euphytica DOI /s z Veitia RA, Bottani S, Birchler JA Gene dosage effects: nonlinearities, genetic interactions, and dosage compensation. Trends in Genetics 29: Wang J, Tian L, Lee HS, Wei NE, Jiang H, Watson B, Madlung A, Osborn TC, Doerge RW, Comai L, Chen ZJ Genomewide nonadditive gene regulation in arabidopsis allotetraploids. Genetics 172: Wang X, Zhang H, Li Y, Zhang Z, Li L, Liu B Transcriptome asymmetry in synthetic and natural allotetraploid wheats, revealed by RNA-sequencing. New Phytologist 209: Yoo MJ, Liu X, Pires JC, Soltis PS, Soltis DE Nonadditive gene expression in polyploids. Annual Review of Genetics 48:
23 Table 1. Determination of the subgenome origin of homoeolog loss (A) and silencing events (B) detected in two accessions of C. arabica. A Acc. of C. arabica Number of homoeolog loss events Subgenome origin of homoeolog losses C a E a Not determined AR Caturra Loss shared by the 2 acc Loss not shared B Acc. of C. arabica Number of silenced genes Subgenome origin of silenced genes C a E a Not determined AR Caturra Silencing shared by the 2 acc Silencing not shared
24 Table 2. Characterization of four regions in C. arabica (acc. Caturra) carrying contiguous genes exhibiting homoeolog losses using the 11 chromosomes of C. canephora as genomic reference sequence. Region code Homoeologous chromosome groups Number of genes exhibiting homoeolog loss Gene identifier (start/end) Size (kb) A 2 4 Cc02g15410/Cc02g B 2 7 Cc02g39930/Cc02g C Cc07g00010/Cc07g D 10 3 Cc10g00010/Cc10g
25 Table 3. Gene ontology enrichment analysis of genes exhibiting homoeolog silencing shared by the three accessions of C. arabica analyzed. Analysis was performed using the full set of 9,047 analyzed genes as reference group and Fisher s exact test with a false discovery rate (FDR) correction for multiple testing. GO-ID Term Category FDR P-Value Test Ref notannottest notannotref Over/Under GO: Intracellular organelle C E E Under GO: Organelle C E E Under GO: Membrane-bounded organelle C E E Under GO: Intracellular membrane-bounded organelle C E E Under GO: Cell C E E Under GO: Cell part C E E Under GO: Intracellular part C E E Under GO: Cellular nitrogen compound metabolic process P E E Under GO: Intracellular C E E Under GO: Cellular aromatic compound metabolic process P E E Under GO: Organic cyclic compound metabolic process P E E Under GO: Heterocycle metabolic process P E E Under GO: Nucleobase-containing compound metabolic p. P E E Under GO: Nucleus C E E Under GO: Cytoplasmic part C E E Under GO: Gene expression P E E Under GO: Nucleic acid binding F E E Under GO: Cellular macromolecule biosynthetic process P E E Under GO: Cellular biosynthetic process P E E Under GO: Macromolecule biosynthetic process P E E Under GO: Cellular nitrogen compound biosynthetic proc. P E E Under GO: Organic substance biosynthetic process P E E Under GO: Nitrogen compound metabolic process P E E Under GO: Cytoplasm C E E Under GO: Cellular metabolic process P E E Under 25
26 Figure 1. Flow chart of methods used to analyze the fate of duplicate genes in allotetraploid C. arabica. 26
27 Figure 2. GC-content normalized DNA copy number profile and FREEC-predicted copy number alteration (Red: gains; blue: losses) for the C. arabica genome (acc. Caturra) using a non-overlapping 50 kb sliding window and the 11 chromosomes of C. canephora as genomic reference sequence. Automatically predicted copy numbers are shown in black (line). 27
28 Figure 3. SNP density along the 11 homoeologous chromosome groups in C. arabica (acc. Caturra). The 11 chromosomes of C. canephora (acc. DH200-94) were used as reference genome. Non-overlapping 10 kb sliding windows and coverage criteria required to consider a position to minimize the rate of false SNPs were applied to estimate the density of SNPs in C. arabica. The relative proportion (percentage nucleotides) in C. canephora (1 Mb sliding window) of transposable elements (green) and genes (blue) are shown at the bottom. 28
29 Figure 4. Examples of regions exhibiting homoeologous SNP deficit in C. arabica (acc. Caturra) on homoeologous chromosome groups 2 and 7. Non-overlapping 10 kb sliding windows were used to estimate SNP density in C. arabica (Yellow track). To minimize the rate of false-positive SNPs, a minimum depth coverage of 10 was required for a position to be taken into consideration, and positions with a depth coverage more than twice the overall sample mean were discarded (Blue track). 29
Reasons for the study
 Systematic study Wittall J.B. et al. (2010): Finding a (pine) needle in a haystack: chloroplast genome sequence divergence in rare and widespread pines. Molecular Ecology 19, 100-114. Reasons for the study
Systematic study Wittall J.B. et al. (2010): Finding a (pine) needle in a haystack: chloroplast genome sequence divergence in rare and widespread pines. Molecular Ecology 19, 100-114. Reasons for the study
WP Board 1054/08 Rev. 1
 WP Board 1054/08 Rev. 1 9 September 2009 Original: English E Executive Board/ International Coffee Council 22 25 September 2009 London, England Sequencing the genome for enhanced characterization, utilization,
WP Board 1054/08 Rev. 1 9 September 2009 Original: English E Executive Board/ International Coffee Council 22 25 September 2009 London, England Sequencing the genome for enhanced characterization, utilization,
Chapter V SUMMARY AND CONCLUSION
 Chapter V SUMMARY AND CONCLUSION Coffea is economically the most important genus of the family Rubiaceae, producing the coffee of commerce. Coffee of commerce is obtained mainly from Coffea arabica and
Chapter V SUMMARY AND CONCLUSION Coffea is economically the most important genus of the family Rubiaceae, producing the coffee of commerce. Coffee of commerce is obtained mainly from Coffea arabica and
Where in the Genome is the Flax b1 Locus?
 Where in the Genome is the Flax b1 Locus? Kayla Lindenback 1 and Helen Booker 2 1,2 Plant Sciences Department, University of Saskatchewan, Saskatoon, SK S7N 5A8 2 Crop Development Center, University of
Where in the Genome is the Flax b1 Locus? Kayla Lindenback 1 and Helen Booker 2 1,2 Plant Sciences Department, University of Saskatchewan, Saskatoon, SK S7N 5A8 2 Crop Development Center, University of
Genome-wide identification and characterization of mirnas responsive to Verticillium longisporum infection in Brassica napus by deep sequencing
 Genome-wide identification and characterization of mirnas responsive to Verticillium longisporum infection in Brassica napus by deep sequencing Longjiang Fan, Dan Shen, Daguang Cai (Zhejiang University/Kiel
Genome-wide identification and characterization of mirnas responsive to Verticillium longisporum infection in Brassica napus by deep sequencing Longjiang Fan, Dan Shen, Daguang Cai (Zhejiang University/Kiel
SNP discovery from amphidiploid species and transferability across the Brassicaceae
 SNP discovery from amphidiploid species and transferability across the Brassicaceae Jacqueline Batley University of Queensland, Australia j.batley@uq.edu.au 1 Outline Objectives Brassicas Genome Sequencing
SNP discovery from amphidiploid species and transferability across the Brassicaceae Jacqueline Batley University of Queensland, Australia j.batley@uq.edu.au 1 Outline Objectives Brassicas Genome Sequencing
MBA 503 Final Project Guidelines and Rubric
 MBA 503 Final Project Guidelines and Rubric Overview There are two summative assessments for this course. For your first assessment, you will be objectively assessed by your completion of a series of MyAccountingLab
MBA 503 Final Project Guidelines and Rubric Overview There are two summative assessments for this course. For your first assessment, you will be objectively assessed by your completion of a series of MyAccountingLab
Genetic diversity of wild Coffee (Coffea arabica) and its implication for conservation
 Genetic diversity of wild Coffee (Coffea arabica) and its implication for conservation Kassahun Tesfaye, Feyera Senbeta, Tamiru Oljira, Solomon Balemi, Govers, K., Endashaw Bekele, Borsch, T. Biodiversity
Genetic diversity of wild Coffee (Coffea arabica) and its implication for conservation Kassahun Tesfaye, Feyera Senbeta, Tamiru Oljira, Solomon Balemi, Govers, K., Endashaw Bekele, Borsch, T. Biodiversity
Catalogue of published works on. Maize Lethal Necrosis (MLN) Disease
 Catalogue of published works on Maize Lethal Necrosis (MLN) Disease Mentions of Maize Lethal Necrosis (MLN) Disease - Reports and Journals Current and future potential distribution of maize chlorotic mottle
Catalogue of published works on Maize Lethal Necrosis (MLN) Disease Mentions of Maize Lethal Necrosis (MLN) Disease - Reports and Journals Current and future potential distribution of maize chlorotic mottle
Gasoline Empirical Analysis: Competition Bureau March 2005
 Gasoline Empirical Analysis: Update of Four Elements of the January 2001 Conference Board study: "The Final Fifteen Feet of Hose: The Canadian Gasoline Industry in the Year 2000" Competition Bureau March
Gasoline Empirical Analysis: Update of Four Elements of the January 2001 Conference Board study: "The Final Fifteen Feet of Hose: The Canadian Gasoline Industry in the Year 2000" Competition Bureau March
Technology: What is in the Sorghum Pipeline
 Technology: What is in the Sorghum Pipeline Zhanguo Xin Gloria Burow Chad Hayes Yves Emendack Lan Liu-Gitz, Halee Hughes, Jacob Sanchez, DeeDee Laumbach, Matt Nesbitt ENVIRONMENTAL CHALLENGES REDUCE YIELDS
Technology: What is in the Sorghum Pipeline Zhanguo Xin Gloria Burow Chad Hayes Yves Emendack Lan Liu-Gitz, Halee Hughes, Jacob Sanchez, DeeDee Laumbach, Matt Nesbitt ENVIRONMENTAL CHALLENGES REDUCE YIELDS
THE MANIFOLD EFFECTS OF GENES AFFECTING FRUIT SIZE AND VEGETATIVE GROWTH IN THE RASPBERRY
 THE MANIFOLD EFFECTS OF GENES AFFECTING FRUIT SIZE AND VEGETATIVE GROWTH IN THE RASPBERRY II. GENE I2 BY D. L. JENNINGS Scottish Horticultural Research Institute, Dundee {Received 16 September 1965)...
THE MANIFOLD EFFECTS OF GENES AFFECTING FRUIT SIZE AND VEGETATIVE GROWTH IN THE RASPBERRY II. GENE I2 BY D. L. JENNINGS Scottish Horticultural Research Institute, Dundee {Received 16 September 1965)...
Online Appendix to. Are Two heads Better Than One: Team versus Individual Play in Signaling Games. David C. Cooper and John H.
 Online Appendix to Are Two heads Better Than One: Team versus Individual Play in Signaling Games David C. Cooper and John H. Kagel This appendix contains a discussion of the robustness of the regression
Online Appendix to Are Two heads Better Than One: Team versus Individual Play in Signaling Games David C. Cooper and John H. Kagel This appendix contains a discussion of the robustness of the regression
MUMmer 2.0. Original implementation required large amounts of memory
 Rationale: MUMmer 2.0 Original implementation required large amounts of memory Advantages: Chromosome scale inversions in bacteria Large scale duplications in Arabidopsis Ancient human duplications when
Rationale: MUMmer 2.0 Original implementation required large amounts of memory Advantages: Chromosome scale inversions in bacteria Large scale duplications in Arabidopsis Ancient human duplications when
Missing value imputation in SAS: an intro to Proc MI and MIANALYZE
 Victoria SAS Users Group November 26, 2013 Missing value imputation in SAS: an intro to Proc MI and MIANALYZE Sylvain Tremblay SAS Canada Education Copyright 2010 SAS Institute Inc. All rights reserved.
Victoria SAS Users Group November 26, 2013 Missing value imputation in SAS: an intro to Proc MI and MIANALYZE Sylvain Tremblay SAS Canada Education Copyright 2010 SAS Institute Inc. All rights reserved.
The poor lonesome A subgenome of Brassica napus var. Darmor (AACC) may not survive without its mate
 Research The poor lonesome A subgenome of Brassica napus var. Darmor (AACC) may not survive without its mate Alexandre Pele*, Gwenn Trotoux*,Frederique Eber, Maryse Lode, Marie Gilet, Gwenaelle Deniot,
Research The poor lonesome A subgenome of Brassica napus var. Darmor (AACC) may not survive without its mate Alexandre Pele*, Gwenn Trotoux*,Frederique Eber, Maryse Lode, Marie Gilet, Gwenaelle Deniot,
COMPARISON OF CORE AND PEEL SAMPLING METHODS FOR DRY MATTER MEASUREMENT IN HASS AVOCADO FRUIT
 New Zealand Avocado Growers' Association Annual Research Report 2004. 4:36 46. COMPARISON OF CORE AND PEEL SAMPLING METHODS FOR DRY MATTER MEASUREMENT IN HASS AVOCADO FRUIT J. MANDEMAKER H. A. PAK T. A.
New Zealand Avocado Growers' Association Annual Research Report 2004. 4:36 46. COMPARISON OF CORE AND PEEL SAMPLING METHODS FOR DRY MATTER MEASUREMENT IN HASS AVOCADO FRUIT J. MANDEMAKER H. A. PAK T. A.
Mapping and Detection of Downy Mildew and Botrytis bunch rot Resistance Loci in Norton-based Population
 Mapping and Detection of Downy Mildew and Botrytis bunch rot Resistance Loci in Norton-based Population Chin-Feng Hwang, Ph.D. State Fruit Experiment Station Darr College of Agriculture Vitis aestivalis-derived
Mapping and Detection of Downy Mildew and Botrytis bunch rot Resistance Loci in Norton-based Population Chin-Feng Hwang, Ph.D. State Fruit Experiment Station Darr College of Agriculture Vitis aestivalis-derived
Supplemental Data. Jeong et al. (2012). Plant Cell /tpc
 Suppmemental Figure 1. Alignment of amino acid sequences of Glycine max JAG1 and its homeolog JAG2, At-JAG and NUBBIN from Arabidopsis thaliana, LYRATE from Solanum lycopersicum, and Zm- JAG from Zea mays.
Suppmemental Figure 1. Alignment of amino acid sequences of Glycine max JAG1 and its homeolog JAG2, At-JAG and NUBBIN from Arabidopsis thaliana, LYRATE from Solanum lycopersicum, and Zm- JAG from Zea mays.
Apport de la Cytogénétique Moléculaire. àl analyse du Génome de la Canne à sucre
 Apport de la Cytogénétique Moléculaire àl analyse du Génome de la Canne à sucre Maguy Rodier, Lolita Triaire, Angélique D Hont in collaboration with BSES, Australia : Nathalie & George Piperidis USP, Brazil
Apport de la Cytogénétique Moléculaire àl analyse du Génome de la Canne à sucre Maguy Rodier, Lolita Triaire, Angélique D Hont in collaboration with BSES, Australia : Nathalie & George Piperidis USP, Brazil
STATE OF THE VITIVINICULTURE WORLD MARKET
 STATE OF THE VITIVINICULTURE WORLD MARKET April 2015 1 Table of contents 1. 2014 VITIVINICULTURAL PRODUCTION POTENTIAL 3 2. WINE PRODUCTION 5 3. WINE CONSUMPTION 7 4. INTERNATIONAL TRADE 9 Abbreviations:
STATE OF THE VITIVINICULTURE WORLD MARKET April 2015 1 Table of contents 1. 2014 VITIVINICULTURAL PRODUCTION POTENTIAL 3 2. WINE PRODUCTION 5 3. WINE CONSUMPTION 7 4. INTERNATIONAL TRADE 9 Abbreviations:
EFFECT OF TOMATO GENETIC VARIATION ON LYE PEELING EFFICACY TOMATO SOLUTIONS JIM AND ADAM DICK SUMMARY
 EFFECT OF TOMATO GENETIC VARIATION ON LYE PEELING EFFICACY TOMATO SOLUTIONS JIM AND ADAM DICK 2013 SUMMARY Several breeding lines and hybrids were peeled in an 18% lye solution using an exposure time of
EFFECT OF TOMATO GENETIC VARIATION ON LYE PEELING EFFICACY TOMATO SOLUTIONS JIM AND ADAM DICK 2013 SUMMARY Several breeding lines and hybrids were peeled in an 18% lye solution using an exposure time of
IT 403 Project Beer Advocate Analysis
 1. Exploratory Data Analysis (EDA) IT 403 Project Beer Advocate Analysis Beer Advocate is a membership-based reviews website where members rank different beers based on a wide number of categories. The
1. Exploratory Data Analysis (EDA) IT 403 Project Beer Advocate Analysis Beer Advocate is a membership-based reviews website where members rank different beers based on a wide number of categories. The
Organization, diversity, expression and evolutionary dynamics of the NB resistance gene family in grapevine and related species
 Organization, diversity, expression and evolutionary dynamics of the NB resistance gene family in grapevine and related species guillaume.barnabe@inra.fr Rustenholz Camille camille.rustenholz@inra.fr Merdinoglu
Organization, diversity, expression and evolutionary dynamics of the NB resistance gene family in grapevine and related species guillaume.barnabe@inra.fr Rustenholz Camille camille.rustenholz@inra.fr Merdinoglu
The Roles of Social Media and Expert Reviews in the Market for High-End Goods: An Example Using Bordeaux and California Wines
 The Roles of Social Media and Expert Reviews in the Market for High-End Goods: An Example Using Bordeaux and California Wines Alex Albright, Stanford/Harvard University Peter Pedroni, Williams College
The Roles of Social Media and Expert Reviews in the Market for High-End Goods: An Example Using Bordeaux and California Wines Alex Albright, Stanford/Harvard University Peter Pedroni, Williams College
Big Data and the Productivity Challenge for Wine Grapes. Nick Dokoozlian Agricultural Outlook Forum February
 Big Data and the Productivity Challenge for Wine Grapes Nick Dokoozlian Agricultural Outlook Forum February 2016 0 Big Data and the Productivity Challenge for Wine Grapes Outline Current production challenges
Big Data and the Productivity Challenge for Wine Grapes Nick Dokoozlian Agricultural Outlook Forum February 2016 0 Big Data and the Productivity Challenge for Wine Grapes Outline Current production challenges
Online Appendix to Voluntary Disclosure and Information Asymmetry: Evidence from the 2005 Securities Offering Reform
 Online Appendix to Voluntary Disclosure and Information Asymmetry: Evidence from the 2005 Securities Offering Reform This document contains several additional results that are untabulated but referenced
Online Appendix to Voluntary Disclosure and Information Asymmetry: Evidence from the 2005 Securities Offering Reform This document contains several additional results that are untabulated but referenced
Using Growing Degree Hours Accumulated Thirty Days after Bloom to Help Growers Predict Difficult Fruit Sizing Years
 Using Growing Degree Hours Accumulated Thirty Days after Bloom to Help Growers Predict Difficult Fruit Sizing Years G. Lopez 1 and T. DeJong 2 1 Àrea de Tecnologia del Reg, IRTA, Lleida, Spain 2 Department
Using Growing Degree Hours Accumulated Thirty Days after Bloom to Help Growers Predict Difficult Fruit Sizing Years G. Lopez 1 and T. DeJong 2 1 Àrea de Tecnologia del Reg, IRTA, Lleida, Spain 2 Department
(Definition modified from APSnet)
 Development of a New Clubroot Differential Set S.E. Strelkov, T. Cao, V.P. Manolii and S.F. Hwang Clubroot Summit Edmonton, March 7, 2012 Background Multiple strains of P. brassicae are known to exist
Development of a New Clubroot Differential Set S.E. Strelkov, T. Cao, V.P. Manolii and S.F. Hwang Clubroot Summit Edmonton, March 7, 2012 Background Multiple strains of P. brassicae are known to exist
Predicting Wine Quality
 March 8, 2016 Ilker Karakasoglu Predicting Wine Quality Problem description: You have been retained as a statistical consultant for a wine co-operative, and have been asked to analyze these data. Each
March 8, 2016 Ilker Karakasoglu Predicting Wine Quality Problem description: You have been retained as a statistical consultant for a wine co-operative, and have been asked to analyze these data. Each
OF THE VARIOUS DECIDUOUS and
 (9) PLAXICO, JAMES S. 1955. PROBLEMS OF FACTOR-PRODUCT AGGRE- GATION IN COBB-DOUGLAS VALUE PRODUCTIVITY ANALYSIS. JOUR. FARM ECON. 37: 644-675, ILLUS. (10) SCHICKELE, RAINER. 1941. EFFECT OF TENURE SYSTEMS
(9) PLAXICO, JAMES S. 1955. PROBLEMS OF FACTOR-PRODUCT AGGRE- GATION IN COBB-DOUGLAS VALUE PRODUCTIVITY ANALYSIS. JOUR. FARM ECON. 37: 644-675, ILLUS. (10) SCHICKELE, RAINER. 1941. EFFECT OF TENURE SYSTEMS
THE EFFECT OF GIRDLING ON FRUIT QUALITY, PHENOLOGY AND MINERAL ANALYSIS OF THE AVOCADO TREE
 California Avocado Society 1971-72 Yearbook 55: 162-169 THE EFFECT OF GIRDLING ON FRUIT QUALITY, PHENOLOGY AND MINERAL ANALYSIS OF THE AVOCADO TREE E. Lahav Division of Subtropical Horticulture, The Volcani
California Avocado Society 1971-72 Yearbook 55: 162-169 THE EFFECT OF GIRDLING ON FRUIT QUALITY, PHENOLOGY AND MINERAL ANALYSIS OF THE AVOCADO TREE E. Lahav Division of Subtropical Horticulture, The Volcani
Reshaping of crossover distribution in Vitis vinifera x Muscadinia rotundifolia interspecific hybrids
 Reshaping of crossover distribution in Vitis vinifera Muscadinia rotundifolia interspecific hybrids Marion Delame, Emilce Prado, Sophie Blanc, Guillaume Robert-Siegwald, Christophe Schneider, Pere Mestre,
Reshaping of crossover distribution in Vitis vinifera Muscadinia rotundifolia interspecific hybrids Marion Delame, Emilce Prado, Sophie Blanc, Guillaume Robert-Siegwald, Christophe Schneider, Pere Mestre,
ICC September 2018 Original: English. Emerging coffee markets: South and East Asia
 ICC 122-6 7 September 2018 Original: English E International Coffee Council 122 st Session 17 21 September 2018 London, UK Emerging coffee markets: South and East Asia Background 1. In accordance with
ICC 122-6 7 September 2018 Original: English E International Coffee Council 122 st Session 17 21 September 2018 London, UK Emerging coffee markets: South and East Asia Background 1. In accordance with
Interloper s legacy: invasive, hybrid-derived California wild radish (Raphanus sativus) evolves to outperform its immigrant parents
 Interloper s legacy: invasive, hybrid-derived California wild radish (Raphanus sativus) evolves to outperform its immigrant parents Caroline E. Ridley 1 and Norman C. Ellstrand 1,2 1 Department of Botany
Interloper s legacy: invasive, hybrid-derived California wild radish (Raphanus sativus) evolves to outperform its immigrant parents Caroline E. Ridley 1 and Norman C. Ellstrand 1,2 1 Department of Botany
SHORT TERM SCIENTIFIC MISSIONS (STSMs)
 SHORT TERM SCIENTIFIC MISSIONS (STSMs) Reference: Short Term Scientific Mission, COST Action FA1003 Beneficiary: Bocharova Valeriia, National Scientific Center Institute of viticulture and winemaking named
SHORT TERM SCIENTIFIC MISSIONS (STSMs) Reference: Short Term Scientific Mission, COST Action FA1003 Beneficiary: Bocharova Valeriia, National Scientific Center Institute of viticulture and winemaking named
Pevzner P., Tesler G. PNAS 2003;100: Copyright 2003, The National Academy of Sciences
 Two different most parsimonious scenarios that transform the order of the 11 synteny blocks on the mouse X chromosome into the order on the human X chromosome Pevzner P., Tesler G. PNAS 2003;100:7672-7677
Two different most parsimonious scenarios that transform the order of the 11 synteny blocks on the mouse X chromosome into the order on the human X chromosome Pevzner P., Tesler G. PNAS 2003;100:7672-7677
Work Sample (Minimum) for 10-K Integration Assignment MAN and for suppliers of raw materials and services that the Company relies on.
 Work Sample (Minimum) for 10-K Integration Assignment MAN 4720 Employee Name: Your name goes here Company: Starbucks Date of Your Report: Date of 10-K: PESTEL 1. Political: Pg. 5 The Company supports the
Work Sample (Minimum) for 10-K Integration Assignment MAN 4720 Employee Name: Your name goes here Company: Starbucks Date of Your Report: Date of 10-K: PESTEL 1. Political: Pg. 5 The Company supports the
Confectionary sunflower A new breeding program. Sun Yue (Jenny)
 Confectionary sunflower A new breeding program Sun Yue (Jenny) Sunflower in Australia Oilseed: vegetable oil, margarine Canola, cotton seeds account for >90% of oilseed production Sunflower less competitive
Confectionary sunflower A new breeding program Sun Yue (Jenny) Sunflower in Australia Oilseed: vegetable oil, margarine Canola, cotton seeds account for >90% of oilseed production Sunflower less competitive
Colorado State University Viticulture and Enology. Grapevine Cold Hardiness
 Colorado State University Viticulture and Enology Grapevine Cold Hardiness Grapevine cold hardiness is dependent on multiple independent variables such as variety and clone, shoot vigor, previous season
Colorado State University Viticulture and Enology Grapevine Cold Hardiness Grapevine cold hardiness is dependent on multiple independent variables such as variety and clone, shoot vigor, previous season
GENETICS AND EVOLUTION OF CORN. This activity previews basic concepts of inheritance and how species change over time.
 GENETICS AND EVOLUTION OF CORN This activity previews basic concepts of inheritance and how species change over time. Objectives for Exam #1: 1. Describe and complete a monohybrid ( one trait ) cross of
GENETICS AND EVOLUTION OF CORN This activity previews basic concepts of inheritance and how species change over time. Objectives for Exam #1: 1. Describe and complete a monohybrid ( one trait ) cross of
Genomics: cracking the mysteries of walnuts
 Review Article Genomics: cracking the mysteries of walnuts Fei Chen 1*#, Junhao Chen 2*, Zhengjia Wang 2, Jiawei Zhang 1, Meigui Lin 1, Liangsheng Zhang 1# 1 State Key Laboratory of Ecological Pest Control
Review Article Genomics: cracking the mysteries of walnuts Fei Chen 1*#, Junhao Chen 2*, Zhengjia Wang 2, Jiawei Zhang 1, Meigui Lin 1, Liangsheng Zhang 1# 1 State Key Laboratory of Ecological Pest Control
THE ROLE OF CAROTENOID CLEAVAGE DIOXYGENASE 4 IN FLOWER COLOR OF THE ALLOPOLYPLOID BRASSICA NAPUS. A Thesis. Presented to
 THE ROLE OF CAROTENOID CLEAVAGE DIOXYGENASE 4 IN FLOWER COLOR OF THE ALLOPOLYPLOID BRASSICA NAPUS A Thesis Presented to The Faculty of California Polytechnic State University, San Luis Obispo In Partial
THE ROLE OF CAROTENOID CLEAVAGE DIOXYGENASE 4 IN FLOWER COLOR OF THE ALLOPOLYPLOID BRASSICA NAPUS A Thesis Presented to The Faculty of California Polytechnic State University, San Luis Obispo In Partial
Biologist at Work! Experiment: Width across knuckles of: left hand. cm... right hand. cm. Analysis: Decision: /13 cm. Name
 wrong 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 right 72 71 70 69 68 67 66 65 64 63 62 61 60 59 58 57 56 55 54 53 52 score 100 98.6 97.2 95.8 94.4 93.1 91.7 90.3 88.9 87.5 86.1 84.7 83.3 81.9
wrong 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 right 72 71 70 69 68 67 66 65 64 63 62 61 60 59 58 57 56 55 54 53 52 score 100 98.6 97.2 95.8 94.4 93.1 91.7 90.3 88.9 87.5 86.1 84.7 83.3 81.9
FINAL REPORT TO AUSTRALIAN GRAPE AND WINE AUTHORITY. Project Number: AGT1524. Principal Investigator: Ana Hranilovic
 Collaboration with Bordeaux researchers to explore genotypic and phenotypic diversity of Lachancea thermotolerans - a promising non- Saccharomyces for winemaking FINAL REPORT TO AUSTRALIAN GRAPE AND WINE
Collaboration with Bordeaux researchers to explore genotypic and phenotypic diversity of Lachancea thermotolerans - a promising non- Saccharomyces for winemaking FINAL REPORT TO AUSTRALIAN GRAPE AND WINE
EFFECTS OF HIGH TEMPERATURE AND CONTROLLED FRUITING ON COTTON YIELD
 Chapter 6 57 EFFECTS OF HIGH TEMPERATURE AND CONTROLLED FRUITING ON COTTON YIELD Carl F. Ehlig USDA-ARS Brawley, California INTRODUCTION The fruit load is the primary cause for mid-season decreases in
Chapter 6 57 EFFECTS OF HIGH TEMPERATURE AND CONTROLLED FRUITING ON COTTON YIELD Carl F. Ehlig USDA-ARS Brawley, California INTRODUCTION The fruit load is the primary cause for mid-season decreases in
Which of your fingernails comes closest to 1 cm in width? What is the length between your thumb tip and extended index finger tip? If no, why not?
 wrong 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 right 66 65 64 63 62 61 60 59 58 57 56 55 54 53 52 51 50 49 score 100 98.5 97.0 95.5 93.9 92.4 90.9 89.4 87.9 86.4 84.8 83.3 81.8 80.3 78.8 77.3 75.8 74.2
wrong 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 right 66 65 64 63 62 61 60 59 58 57 56 55 54 53 52 51 50 49 score 100 98.5 97.0 95.5 93.9 92.4 90.9 89.4 87.9 86.4 84.8 83.3 81.8 80.3 78.8 77.3 75.8 74.2
Napa County Planning Commission Board Agenda Letter
 Agenda Date: 7/1/2015 Agenda Placement: 10A Continued From: May 20, 2015 Napa County Planning Commission Board Agenda Letter TO: FROM: Napa County Planning Commission John McDowell for David Morrison -
Agenda Date: 7/1/2015 Agenda Placement: 10A Continued From: May 20, 2015 Napa County Planning Commission Board Agenda Letter TO: FROM: Napa County Planning Commission John McDowell for David Morrison -
EFFECT OF MODE OF RIPENING ON ETHYLENE BIOSYNTHESIS DURING RIPENING OF ONE DIPLOID BANANA FRUIT
 EFFECT OF MODE OF RIPENING ON ETHYLENE BIOSYNTHESIS DURING RIPENING OF ONE DIPLOID BANANA FRUIT HUBERT O., CHILLET M., JULIANNUS P., FILS-LYCAON B., MBEGUIE-A-MBEGUIE* D. * CIRAD/UMR 94 QUALITROP, Neufchâteau,
EFFECT OF MODE OF RIPENING ON ETHYLENE BIOSYNTHESIS DURING RIPENING OF ONE DIPLOID BANANA FRUIT HUBERT O., CHILLET M., JULIANNUS P., FILS-LYCAON B., MBEGUIE-A-MBEGUIE* D. * CIRAD/UMR 94 QUALITROP, Neufchâteau,
COOPER COMPARISONS Next Phase of Study: Results with Wine
 COOPER COMPARISONS Next Phase of Study: Results with Wine A follow-up study has just been completed, with the generous cooperation of Cakebread Cellars, Lafond Winery, and Edna Valley Vineyards. Many of
COOPER COMPARISONS Next Phase of Study: Results with Wine A follow-up study has just been completed, with the generous cooperation of Cakebread Cellars, Lafond Winery, and Edna Valley Vineyards. Many of
Structures of Life. Investigation 1: Origin of Seeds. Big Question: 3 rd Science Notebook. Name:
 3 rd Science Notebook Structures of Life Investigation 1: Origin of Seeds Name: Big Question: What are the properties of seeds and how does water affect them? 1 Alignment with New York State Science Standards
3 rd Science Notebook Structures of Life Investigation 1: Origin of Seeds Name: Big Question: What are the properties of seeds and how does water affect them? 1 Alignment with New York State Science Standards
Relation between Grape Wine Quality and Related Physicochemical Indexes
 Research Journal of Applied Sciences, Engineering and Technology 5(4): 557-5577, 013 ISSN: 040-7459; e-issn: 040-7467 Maxwell Scientific Organization, 013 Submitted: October 1, 01 Accepted: December 03,
Research Journal of Applied Sciences, Engineering and Technology 5(4): 557-5577, 013 ISSN: 040-7459; e-issn: 040-7467 Maxwell Scientific Organization, 013 Submitted: October 1, 01 Accepted: December 03,
INVESTIGATIONS INTO THE RELATIONSHIPS OF STRESS AND LEAF HEALTH OF THE GRAPEVINE (VITIS VINIFERA L.) ON GRAPE AND WINE QUALITIES
 INVESTIGATIONS INTO THE RELATIONSHIPS OF STRESS AND LEAF HEALTH OF THE GRAPEVINE (VITIS VINIFERA L.) ON GRAPE AND WINE QUALITIES by Reuben Wells BAgrSc (Hons) Submitted in fulfilment of the requirements
INVESTIGATIONS INTO THE RELATIONSHIPS OF STRESS AND LEAF HEALTH OF THE GRAPEVINE (VITIS VINIFERA L.) ON GRAPE AND WINE QUALITIES by Reuben Wells BAgrSc (Hons) Submitted in fulfilment of the requirements
Laboratory Performance Assessment. Report. Analysis of Pesticides and Anthraquinone. in Black Tea
 Laboratory Performance Assessment Report Analysis of Pesticides and Anthraquinone in Black Tea May 2013 Summary This laboratory performance assessment on pesticides in black tea was designed and organised
Laboratory Performance Assessment Report Analysis of Pesticides and Anthraquinone in Black Tea May 2013 Summary This laboratory performance assessment on pesticides in black tea was designed and organised
QUALITY, PRICING AND THE PERFORMANCE OF THE WHEAT INDUSTRY IN SOUTH AFRICA
 QUALITY, PRICING AND THE PERFORMANCE OF THE WHEAT INDUSTRY IN SOUTH AFRICA 21 September 2015 Dr Johnny van der Merwe Lecturer / Agricultural economics (Prof HD van Schalkwyk and Dr PC Cloete) So what motivated
QUALITY, PRICING AND THE PERFORMANCE OF THE WHEAT INDUSTRY IN SOUTH AFRICA 21 September 2015 Dr Johnny van der Merwe Lecturer / Agricultural economics (Prof HD van Schalkwyk and Dr PC Cloete) So what motivated
CHAPTER 8. Sample Laboratory Experiments
 CHAPTER 8 Sample Laboratory Experiments 8.a Analytical Experiments without an External Reference Standard; Conformational Identification without Quantification. Jake Ginsbach CAUTION: Do not repeat this
CHAPTER 8 Sample Laboratory Experiments 8.a Analytical Experiments without an External Reference Standard; Conformational Identification without Quantification. Jake Ginsbach CAUTION: Do not repeat this
FACTORS DETERMINING UNITED STATES IMPORTS OF COFFEE
 12 November 1953 FACTORS DETERMINING UNITED STATES IMPORTS OF COFFEE The present paper is the first in a series which will offer analyses of the factors that account for the imports into the United States
12 November 1953 FACTORS DETERMINING UNITED STATES IMPORTS OF COFFEE The present paper is the first in a series which will offer analyses of the factors that account for the imports into the United States
ARM4 Advances: Genetic Algorithm Improvements. Ed Downs & Gianluca Paganoni
 ARM4 Advances: Genetic Algorithm Improvements Ed Downs & Gianluca Paganoni Artificial Intelligence In Trading, we want to identify trades that generate the most consistent profits over a long period of
ARM4 Advances: Genetic Algorithm Improvements Ed Downs & Gianluca Paganoni Artificial Intelligence In Trading, we want to identify trades that generate the most consistent profits over a long period of
Alcoholic Fermentation in Yeast A Bioengineering Design Challenge 1
 Alcoholic Fermentation in Yeast A Bioengineering Design Challenge 1 I. Introduction Yeasts are single cell fungi. People use yeast to make bread, wine and beer. For your experiment, you will use the little
Alcoholic Fermentation in Yeast A Bioengineering Design Challenge 1 I. Introduction Yeasts are single cell fungi. People use yeast to make bread, wine and beer. For your experiment, you will use the little
-SQA- SCOTTISH QUALIFICATIONS AUTHORITY NATIONAL CERTIFICATE MODULE: UNIT SPECIFICATION GENERAL INFORMATION. -Module Number Session
 -SQA- SCOTTISH QUALIFICATIONS AUTHORITY NATIONAL CERTIFICATE MODULE: UNIT SPECIFICATION GENERAL INFORMATION -Module Number- 3230006 -Session-1996-97 -Superclass- NE -Title- CAKE DECORATION: ADVANCED ROYAL
-SQA- SCOTTISH QUALIFICATIONS AUTHORITY NATIONAL CERTIFICATE MODULE: UNIT SPECIFICATION GENERAL INFORMATION -Module Number- 3230006 -Session-1996-97 -Superclass- NE -Title- CAKE DECORATION: ADVANCED ROYAL
Rapid Analysis of Soft Drinks Using the ACQUITY UPLC H-Class System with the Waters Beverage Analysis Kit
 Rapid Analysis of Soft Drinks Using the ACQUITY UPLC H-Class System with the Waters Beverage Analysis Kit Mark E. Benvenuti, Raymond Giska, and Jennifer A. Burgess Waters Corporation, Milford, MA U.S.
Rapid Analysis of Soft Drinks Using the ACQUITY UPLC H-Class System with the Waters Beverage Analysis Kit Mark E. Benvenuti, Raymond Giska, and Jennifer A. Burgess Waters Corporation, Milford, MA U.S.
Title: Development of Simple Sequence Repeat DNA markers for Muscadine Grape Cultivar Identification.
 Title: Development of Simple Sequence Repeat DNA markers for Muscadine Grape Cultivar Identification. Progress Report Grant Code: SRSFC Project # 2018 R-06 Research Proposal Name, Mailing and Email Address
Title: Development of Simple Sequence Repeat DNA markers for Muscadine Grape Cultivar Identification. Progress Report Grant Code: SRSFC Project # 2018 R-06 Research Proposal Name, Mailing and Email Address
Thermal Hydraulic Analysis of 49-2 Swimming Pool Reactor with a. Passive Siphon Breaker
 Thermal Hydraulic Analysis of 49-2 Swimming Pool Reactor with a Passive Siphon Breaker Zhiting Yue 1, Songtao Ji 1 1) China Institute of Atomic Energy(CIAE), Beijing 102413, China Corresponding author:
Thermal Hydraulic Analysis of 49-2 Swimming Pool Reactor with a Passive Siphon Breaker Zhiting Yue 1, Songtao Ji 1 1) China Institute of Atomic Energy(CIAE), Beijing 102413, China Corresponding author:
of Vitis vinifera using
 Characterisation of the pan-genome of Vitis vinifera using Next Generation Sequencing Plant Biology Europe 2018 - June 18-21 - Copenhagen Gabriele Magris (gmagris@appliedgenomics.org) Genetic variation
Characterisation of the pan-genome of Vitis vinifera using Next Generation Sequencing Plant Biology Europe 2018 - June 18-21 - Copenhagen Gabriele Magris (gmagris@appliedgenomics.org) Genetic variation
5. Supporting documents to be provided by the applicant IMPORTANT DISCLAIMER
 Guidance notes on the classification of a flavouring substance with modifying properties and a flavour enhancer 27.5.2014 Contents 1. Purpose 2. Flavouring substances with modifying properties 3. Flavour
Guidance notes on the classification of a flavouring substance with modifying properties and a flavour enhancer 27.5.2014 Contents 1. Purpose 2. Flavouring substances with modifying properties 3. Flavour
Wine-Tasting by Numbers: Using Binary Logistic Regression to Reveal the Preferences of Experts
 Wine-Tasting by Numbers: Using Binary Logistic Regression to Reveal the Preferences of Experts When you need to understand situations that seem to defy data analysis, you may be able to use techniques
Wine-Tasting by Numbers: Using Binary Logistic Regression to Reveal the Preferences of Experts When you need to understand situations that seem to defy data analysis, you may be able to use techniques
Emerging Local Food Systems in the Caribbean and Southern USA July 6, 2014
 Consumers attitudes toward consumption of two different types of juice beverages based on country of origin (local vs. imported) Presented at Emerging Local Food Systems in the Caribbean and Southern USA
Consumers attitudes toward consumption of two different types of juice beverages based on country of origin (local vs. imported) Presented at Emerging Local Food Systems in the Caribbean and Southern USA
Business opportunities and challenges of mainstreaming biodiversity into the agricultural sector
 Business opportunities and challenges of mainstreaming biodiversity into the agricultural sector Mainstreaming biodiversity into the agricultural sector what does this mean? Cultural service Regulating
Business opportunities and challenges of mainstreaming biodiversity into the agricultural sector Mainstreaming biodiversity into the agricultural sector what does this mean? Cultural service Regulating
An Overview of the U.S. Bell Pepper Industry. Trina Biswas, Zhengfei Guan, 1 Feng Wu University of Florida
 An Overview of the U.S. Bell Pepper Industry Trina Biswas, Zhengfei Guan, 1 Feng Wu University of Florida Bell pepper is one of the most widely cultivated vegetable crops in the world. Characterized by
An Overview of the U.S. Bell Pepper Industry Trina Biswas, Zhengfei Guan, 1 Feng Wu University of Florida Bell pepper is one of the most widely cultivated vegetable crops in the world. Characterized by
Wine Australia Wine.com Data Report. July 21, 2017
 Wine Australia Wine.com Data Report July 21, 2017 INTRODUCTION Wine Opinions is a wine market research company focusing on the attitudes, behaviors, and taste preferences of U.S. wine drinkers. Wine Opinions
Wine Australia Wine.com Data Report July 21, 2017 INTRODUCTION Wine Opinions is a wine market research company focusing on the attitudes, behaviors, and taste preferences of U.S. wine drinkers. Wine Opinions
Buying Filberts On a Sample Basis
 E 55 m ^7q Buying Filberts On a Sample Basis Special Report 279 September 1969 Cooperative Extension Service c, 789/0 ite IP") 0, i mi 1910 S R e, `g,,ttsoliktill:torvti EARs srin ITQ, E,6
E 55 m ^7q Buying Filberts On a Sample Basis Special Report 279 September 1969 Cooperative Extension Service c, 789/0 ite IP") 0, i mi 1910 S R e, `g,,ttsoliktill:torvti EARs srin ITQ, E,6
International Journal of Business and Commerce Vol. 3, No.8: Apr 2014[01-10] (ISSN: )
![International Journal of Business and Commerce Vol. 3, No.8: Apr 2014[01-10] (ISSN: ) International Journal of Business and Commerce Vol. 3, No.8: Apr 2014[01-10] (ISSN: )](/thumbs/86/94457514.jpg) The Comparative Influences of Relationship Marketing, National Cultural values, and Consumer values on Consumer Satisfaction between Local and Global Coffee Shop Brands Yi Hsu Corresponding author: Associate
The Comparative Influences of Relationship Marketing, National Cultural values, and Consumer values on Consumer Satisfaction between Local and Global Coffee Shop Brands Yi Hsu Corresponding author: Associate
Title: Genetic Variation of Crabapples ( Malus spp.) found on Governors Island and NYC Area
 Title: Genetic Variation of Crabapples ( Malus spp.) found on Governors Island and NYC Area Team Members: Jianri Chen, Zinan Ma, Iulius Sergiu Moldovan and Xuanzhi Zhao Sponsoring Teacher: Alfred Lwin
Title: Genetic Variation of Crabapples ( Malus spp.) found on Governors Island and NYC Area Team Members: Jianri Chen, Zinan Ma, Iulius Sergiu Moldovan and Xuanzhi Zhao Sponsoring Teacher: Alfred Lwin
NEW ZEALAND WINE FOOD BILL ORAL SUBMISSION OF NEW ZEALAND WINEGROWERS 23 SEPTEMBER Introduction
 NEW ZEALAND WINE PURE DISCOVERY FOOD BILL ORAL SUBMISSION OF NEW ZEALAND WINEGROWERS 23 SEPTEMBER 2010 Introduction 1. New Zealand Winegrowers (NZW) is the national industry organisation representing the
NEW ZEALAND WINE PURE DISCOVERY FOOD BILL ORAL SUBMISSION OF NEW ZEALAND WINEGROWERS 23 SEPTEMBER 2010 Introduction 1. New Zealand Winegrowers (NZW) is the national industry organisation representing the
Level 3 Biology, 2016
 91605 916050 3SUPERVISOR S Level 3 Biology, 2016 91605 Demonstrate understanding of evolutionary processes leading to speciation 2.00 p.m. Thursday 10 November 2016 Credits: Four Achievement Achievement
91605 916050 3SUPERVISOR S Level 3 Biology, 2016 91605 Demonstrate understanding of evolutionary processes leading to speciation 2.00 p.m. Thursday 10 November 2016 Credits: Four Achievement Achievement
STRUCTURES OF PURINES. Uric acid
 INTRODUCTION PURINES Methylxanthines and methyluric acids are secondary plant metabolites derived from purine nucleotides. The most well known methylxanthines are caffeine (1,3,7- trimethylxanthine) and
INTRODUCTION PURINES Methylxanthines and methyluric acids are secondary plant metabolites derived from purine nucleotides. The most well known methylxanthines are caffeine (1,3,7- trimethylxanthine) and
GENOTYPIC AND ENVIRONMENTAL EFFECTS ON BREAD-MAKING QUALITY OF WINTER WHEAT IN ROMANIA
 GENOTYPIC AND ENVIRONMENTAL EFFECTS ON BREAD-MAKING QUALITY OF WINTER WHEAT IN ROMANIA Mihaela Tianu, Nicolae N. Sãulescu and Gheorghe Ittu ABSTRACT Bread-making quality was analysed in two sets of wheat
GENOTYPIC AND ENVIRONMENTAL EFFECTS ON BREAD-MAKING QUALITY OF WINTER WHEAT IN ROMANIA Mihaela Tianu, Nicolae N. Sãulescu and Gheorghe Ittu ABSTRACT Bread-making quality was analysed in two sets of wheat
Peach and Nectarine Cork Spot: A Review of the 1998 Season
 Peach and Nectarine Cork Spot: A Review of the 1998 Season Kevin R. Day Tree Fruit Farm Advisor Tulare County University of California Cooperative Extension Along with many other problems, fruit corking
Peach and Nectarine Cork Spot: A Review of the 1998 Season Kevin R. Day Tree Fruit Farm Advisor Tulare County University of California Cooperative Extension Along with many other problems, fruit corking
Preliminary observation on a spontaneous tricotyledonous mutant in sunflower
 Preliminary observation on a spontaneous tricotyledonous mutant in sunflower Jinguo Hu 1, Jerry F. Miller 1, Junfang Chen 2, Brady A. Vick 1 1 USDA, Agricultural Research Service, Northern Crop Science
Preliminary observation on a spontaneous tricotyledonous mutant in sunflower Jinguo Hu 1, Jerry F. Miller 1, Junfang Chen 2, Brady A. Vick 1 1 USDA, Agricultural Research Service, Northern Crop Science
Identification of haplotypes controlling seedless by genome resequencing of grape
 Identification of haplotypes controlling seedless by genome resequencing of grape Soon-Chun Jeong scjeong@kribb.re.kr Korea Research Institute of Bioscience and Biotechnology Why seedless grape research
Identification of haplotypes controlling seedless by genome resequencing of grape Soon-Chun Jeong scjeong@kribb.re.kr Korea Research Institute of Bioscience and Biotechnology Why seedless grape research
Calvin Lietzow and James Nienhuis Department of Horticulture, University of Wisconsin, 1575 Linden Dr., Madison, WI 53706
 Precocious Yellow Rind Color in Cucurbita moschata Calvin Lietzow and James Nienhuis Department of Horticulture, University of Wisconsin, 1575 Linden Dr., Madison, WI 53706 Amber DeLong and Linda Wessel-Beaver
Precocious Yellow Rind Color in Cucurbita moschata Calvin Lietzow and James Nienhuis Department of Horticulture, University of Wisconsin, 1575 Linden Dr., Madison, WI 53706 Amber DeLong and Linda Wessel-Beaver
THE NATURAL SUSCEPTIBILITY AND ARTIFICIALLY INDUCED FRUIT CRACKING OF SOUR CHERRY CULTIVARS
 THE NATURAL SUSCEPTIBILITY AND ARTIFICIALLY INDUCED FRUIT CRACKING OF SOUR CHERRY CULTIVARS S. Budan Research Institute for Fruit Growing, Pitesti, Romania sergiu_budan@yahoo.com GENERALITIES It is agreed
THE NATURAL SUSCEPTIBILITY AND ARTIFICIALLY INDUCED FRUIT CRACKING OF SOUR CHERRY CULTIVARS S. Budan Research Institute for Fruit Growing, Pitesti, Romania sergiu_budan@yahoo.com GENERALITIES It is agreed
Determination of Caffeine in Coffee Products According to DIN 20481
 Deteration of Caffeine in Coffee Products According to DI 81 Application ote Food Testing & Agriculture Food Authenticity Author Edgar aegele Agilent Technologies, Inc. Waldbronn, Germany Abstract This
Deteration of Caffeine in Coffee Products According to DI 81 Application ote Food Testing & Agriculture Food Authenticity Author Edgar aegele Agilent Technologies, Inc. Waldbronn, Germany Abstract This
ICC July 2010 Original: French. Study. International Coffee Council 105 th Session September 2010 London, England
 ICC 15-2 12 July 21 Original: French Study E International Coffee Council 15 th Session 22 24 September 21 London, England Relations between coffee stocks and prices Background In the context of its programme
ICC 15-2 12 July 21 Original: French Study E International Coffee Council 15 th Session 22 24 September 21 London, England Relations between coffee stocks and prices Background In the context of its programme
Food and beverage services statistics - NACE Rev. 2
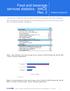 Food and beverage services statistics - NACE Rev. 2 Statistics Explained Data extracted in October 2015. Most recent data: Further Eurostat information, Main tables and Database. This article presents
Food and beverage services statistics - NACE Rev. 2 Statistics Explained Data extracted in October 2015. Most recent data: Further Eurostat information, Main tables and Database. This article presents
IMSI Annual Business Meeting Amherst, Massachusetts October 26, 2008
 Consumer Research to Support a Standardized Grading System for Pure Maple Syrup Presented to: IMSI Annual Business Meeting Amherst, Massachusetts October 26, 2008 Objectives The objectives for the study
Consumer Research to Support a Standardized Grading System for Pure Maple Syrup Presented to: IMSI Annual Business Meeting Amherst, Massachusetts October 26, 2008 Objectives The objectives for the study
RELATIVE EFFICIENCY OF ESTIMATES BASED ON PERCENTAGES OF MISSINGNESS USING THREE IMPUTATION NUMBERS IN MULTIPLE IMPUTATION ANALYSIS ABSTRACT
 RELATIVE EFFICIENCY OF ESTIMATES BASED ON PERCENTAGES OF MISSINGNESS USING THREE IMPUTATION NUMBERS IN MULTIPLE IMPUTATION ANALYSIS Nwakuya, M. T. (Ph.D) Department of Mathematics/Statistics University
RELATIVE EFFICIENCY OF ESTIMATES BASED ON PERCENTAGES OF MISSINGNESS USING THREE IMPUTATION NUMBERS IN MULTIPLE IMPUTATION ANALYSIS Nwakuya, M. T. (Ph.D) Department of Mathematics/Statistics University
Can You Tell the Difference? A Study on the Preference of Bottled Water. [Anonymous Name 1], [Anonymous Name 2]
![Can You Tell the Difference? A Study on the Preference of Bottled Water. [Anonymous Name 1], [Anonymous Name 2] Can You Tell the Difference? A Study on the Preference of Bottled Water. [Anonymous Name 1], [Anonymous Name 2]](/thumbs/78/76759741.jpg) Can You Tell the Difference? A Study on the Preference of Bottled Water [Anonymous Name 1], [Anonymous Name 2] Abstract Our study aims to discover if people will rate the taste of bottled water differently
Can You Tell the Difference? A Study on the Preference of Bottled Water [Anonymous Name 1], [Anonymous Name 2] Abstract Our study aims to discover if people will rate the taste of bottled water differently
INFLUENCE OF ENVIRONMENT - Wine evaporation from barrels By Richard M. Blazer, Enologist Sterling Vineyards Calistoga, CA
 INFLUENCE OF ENVIRONMENT - Wine evaporation from barrels By Richard M. Blazer, Enologist Sterling Vineyards Calistoga, CA Sterling Vineyards stores barrels of wine in both an air-conditioned, unheated,
INFLUENCE OF ENVIRONMENT - Wine evaporation from barrels By Richard M. Blazer, Enologist Sterling Vineyards Calistoga, CA Sterling Vineyards stores barrels of wine in both an air-conditioned, unheated,
Paper Reference IT Principal Learning Information Technology. Level 3 Unit 2: Understanding Organisations
 Centre No. Candidate No. Surname Signature Paper Reference(s) IT302/01 Edexcel Principal Learning Information Technology Level 3 Unit 2: Understanding Organisations Wednesday 3 June 2009 Morning Time:
Centre No. Candidate No. Surname Signature Paper Reference(s) IT302/01 Edexcel Principal Learning Information Technology Level 3 Unit 2: Understanding Organisations Wednesday 3 June 2009 Morning Time:
Is Fair Trade Fair? ARKANSAS C3 TEACHERS HUB. 9-12th Grade Economics Inquiry. Supporting Questions
 9-12th Grade Economics Inquiry Is Fair Trade Fair? Public Domain Image Supporting Questions 1. What is fair trade? 2. If fair trade is so unique, what is free trade? 3. What are the costs and benefits
9-12th Grade Economics Inquiry Is Fair Trade Fair? Public Domain Image Supporting Questions 1. What is fair trade? 2. If fair trade is so unique, what is free trade? 3. What are the costs and benefits
Learning Connectivity Networks from High-Dimensional Point Processes
 Learning Connectivity Networks from High-Dimensional Point Processes Ali Shojaie Department of Biostatistics University of Washington faculty.washington.edu/ashojaie Feb 21st 2018 Motivation: Unlocking
Learning Connectivity Networks from High-Dimensional Point Processes Ali Shojaie Department of Biostatistics University of Washington faculty.washington.edu/ashojaie Feb 21st 2018 Motivation: Unlocking
Wideband HF Channel Availability Measurement Techniques and Results W.N. Furman, J.W. Nieto, W.M. Batts
 Wideband HF Channel Availability Measurement Techniques and Results W.N. Furman, J.W. Nieto, W.M. Batts THIS INFORMATION IS NOT EXPORT CONTROLLED THIS INFORMATION IS APPROVED FOR RELEASE WITHOUT EXPORT
Wideband HF Channel Availability Measurement Techniques and Results W.N. Furman, J.W. Nieto, W.M. Batts THIS INFORMATION IS NOT EXPORT CONTROLLED THIS INFORMATION IS APPROVED FOR RELEASE WITHOUT EXPORT
New from Packaged Facts!
 New from Packaged Facts! FOODSERVICE MARKET INSIGHTS A fresh perspective on the foodservice marketplace Essential Insights on Consumer customerservice@packagedfacts.com (800) 298-5294 (240) 747-3095 (Intl.)
New from Packaged Facts! FOODSERVICE MARKET INSIGHTS A fresh perspective on the foodservice marketplace Essential Insights on Consumer customerservice@packagedfacts.com (800) 298-5294 (240) 747-3095 (Intl.)
DEVELOPMENT OF A RAPID METHOD FOR THE ASSESSMENT OF PHENOLIC MATURITY IN BURGUNDY PINOT NOIR
 PINOT NOIR, PAGE 1 DEVELOPMENT OF A RAPID METHOD FOR THE ASSESSMENT OF PHENOLIC MATURITY IN BURGUNDY PINOT NOIR Eric GRANDJEAN, Centre Œnologique de Bourgogne (COEB)* Christine MONAMY, Bureau Interprofessionnel
PINOT NOIR, PAGE 1 DEVELOPMENT OF A RAPID METHOD FOR THE ASSESSMENT OF PHENOLIC MATURITY IN BURGUNDY PINOT NOIR Eric GRANDJEAN, Centre Œnologique de Bourgogne (COEB)* Christine MONAMY, Bureau Interprofessionnel
Classification Lab (Jelli bellicus) Lab; SB3 b,c
 Classification Lab (Jelli bellicus) Lab; SB3 b,c A branch of biology called taxonomy involves the identification, naming, and classification of species. Assigning scientific names to species is an important
Classification Lab (Jelli bellicus) Lab; SB3 b,c A branch of biology called taxonomy involves the identification, naming, and classification of species. Assigning scientific names to species is an important
2017 FINANCIAL REVIEW
 2017 FINANCIAL REVIEW In addition to activity, strategy, goals, and challenges, survey respondents also provided financial information from 2014, 2015, and 2016. Select results are provided below: 2016
2017 FINANCIAL REVIEW In addition to activity, strategy, goals, and challenges, survey respondents also provided financial information from 2014, 2015, and 2016. Select results are provided below: 2016
Use of a CEP. CEP: What does it mean? Pascale Poukens-Renwart. Certification of Substances Department, EDQM
 Use of a CEP Pascale Poukens-Renwart Certification of Substances Department, EDQM CEP: What does it mean? A chemical or a herbal CEP certifies that the quality of the substance is suitably controlled by
Use of a CEP Pascale Poukens-Renwart Certification of Substances Department, EDQM CEP: What does it mean? A chemical or a herbal CEP certifies that the quality of the substance is suitably controlled by
Detecting Melamine Adulteration in Milk Powder
 Detecting Melamine Adulteration in Milk Powder Introduction Food adulteration is at the top of the list when it comes to food safety concerns, especially following recent incidents, such as the 2008 Chinese
Detecting Melamine Adulteration in Milk Powder Introduction Food adulteration is at the top of the list when it comes to food safety concerns, especially following recent incidents, such as the 2008 Chinese
GLOSSARY Last Updated: 10/17/ KL. Terms and Definitions
 GLOSSARY Last Updated: 10/17/2017 - KL Terms and Definitions Spacing 4ETa Zone(s) Background Drill Elevation Climate Soil Ecoregion 4 Recommended base spacing between containerized, cutting, plug or sprig
GLOSSARY Last Updated: 10/17/2017 - KL Terms and Definitions Spacing 4ETa Zone(s) Background Drill Elevation Climate Soil Ecoregion 4 Recommended base spacing between containerized, cutting, plug or sprig
